[MCWG] Molecular Connectivity Newsletter: May 2026

Thank you to everyone who joined us for April’s MCOS with Dr. Prashin Unadkat! Missed it? The recording is available here.
Date: May 22nd, 2026
Time: 13:00 UTC
Registration: Please register here.
Title: Deciphering the biological correlate of metabolic connectivity in neurodegenerative diseases

Speaker: Dr. Johannes Gnörich is a clinician scientist at the Department of Nuclear Medicine, LMU University Hospital, LMU Munich, Germany. His research focuses on PET imaging, neuroinflammation, and biomarker development in neurodegenerative diseases. Since his doctoral thesis, he has published several translational studies, including methodological evaluations of single-cell radiotracer detection after in vivo tracer injection (scRadiotracing), aiming to disentangle the cellular sources of PET signals in different inflammatory disorders, including neurodegeneration. These studies contributed to the validation and establishment of novel PET tracers in the field. In addition, Dr. Gnörich integrates advanced network-based imaging analyses, particularly metabolic connectivity approaches, to characterize system-level alterations in brain diseases. Moreover, Dr. Gnörich has a strong record of coordinating academic research projects, including recent multi-center studies. He is actively engaged in international collaborations, serves as a peer reviewer for leading scientific journals, and mentors early-career researchers.
Abstract: Metabolic connectivity derived from [18F]FDG PET is an emerging network-based biomarker in neurodegenerative diseases, yet its underlying biological basis remains insufficiently understood. In this talk, I will present our recent published data combining [18F]FDG-PET with scRadiotracing, demonstrating that metabolic connectivity is strongly influenced by cell-specific glucose uptake patterns of neurons, astrocytes, and microglia. Our findings identify astroglial metabolism as a key driver of age-related network reorganization and show that microglial presence and activation states substantially modulate cerebral metabolic networks in translational mouse models. In addition, I will present our ongoing research and preliminary data using single-cell transcriptomics to further dissect the molecular and subcellular mechanisms underlying metabolic connectivity. Together, these approaches aim to establish metabolic connectivity as a biologically informed imaging marker for neurodegenerative disease.
The MCOS promotes rigor in research and resource sharing. We aim to hold MCOS every third Friday of the month, subject to change due to speaker availability. Please stay tuned for MCOS updates and reminders on social media! Thank you!
Each month, we will feature a member of the MCWG and have a brief Q&A!
This month please enjoy our highlight of Prof Dr Christian Habeck, member of the MCWG Steering Committee.

Prof Dr Christian G Habeck studied Physics and after 3 years of undergraduate study at Siegen University in Germany, went to the UK. He did a Master’s Degree in Elementary Particle Theory at the University of Durham, UK, and finally ended up in a DPhil program at the University of Sussex and the Institut Laue Langevin (ILL, Grenoble), studying neutron decay. The Cold War was barely over.
He realized compared to others he saw at ILL, he was not a gifted experimentalist, and at that time the Life Sciences were a promising new venue for Physicists, so he migrated over. He did a Postdoc in Computational Neuroscience at the Neurosciences Institute in San Diego (now closed), before joining Columbia University.
Prof Dr Christian Habeck has graciously responded to our feature questionnaire:
What sparked your interest in molecular imaging or led you to focus on research in molecular imaging?
Arguably, I have not done much molecular imaging at all. But I like being near people who do it for real! I have dealt with 15O and FDG PET data in the olden days… Recently, I have played with the Monash f-PET data set.
What is your role in the Molecular Connectivity Working Group, and what have you been contributing to or working on within the group?
I see myself in contributing to didactic and methodological debates about connectivity analysis, considering statistical robustness and out-of-sample replication; there are lot of people in MCWG with intricate domain knowledge that I don’t have, but what I do have is a lot of experience of playing with fMRI (and formerly PET) data for group-level covariance analysis.
In what ways do you imagine molecular connectivity will advance our understanding of brain function?
Let me phrase it in terms of the differential impact when comparing MC to resting-state functional connectivity (rs-FC): because of better biological grounding with specific tracers or interventions, I think MC can provide more biological constraints than rs-FC and avoid the tendencies of proliferating levels of analytic complexity. I think the achievements of MCWG to date speak for themselves.
What do you think are the most important challenges in current brain connectivity research, or which unsolved/underappreciated issues should the community address?
I think there are some challenges, specifically: temptations we should try to avoid.
(1) Neuroimaging analytics has become overly derivative, a kind of liberal arts with computers, mainly for intellectual ownership. It’s impossible for one person to have their finger on the pulse of a whole field’s Zeitgeist, and these tendencies are fortunately waning it seems, but in my anecdotal experience as a grant reviewer I have seen a lot of complicated Deep-Learning and AI analytics without any grounding in mechanisms, or even specific ideas about which modalities are important etc. Apart from the fact that neuroimaging science does not ONLY consist of outcome prediction and concerns the discovery of mechanisms in large part too, even on the narrow brief of outcome prediction, the superiority of these techniques is often taken as a given, judging from the absence of validation plans in comparison to simple models using first-order moments and demographics. I find this alarming on several levels. I think we should know better than to fall prey to the delusion that adding lots of, often WIDE, data will by itself result in more robust models and that concerns like over-fitting and out-of-sample replication are somehow obviated by deep learning architectures with lots of parameters. If we cannot be clear-eyed about this, I am afraid for society at large when ideas of “predictive policing” and other AI-predictive analytics are pushed hard onto the public.
(2) Further, I would say the field cognitive neuroscience should reverse its move away from cognitive-task imaging for the convenience of resting-BOLD. Tasks can be designed to be rich and ecologically valid; they don’t have to be constrained to bare “lab classics” like the Sternberg working-memory task from the 1960s, but imaging people while performing a task cannot be easily beaten by just recording spontaneous unconstrained activity. We need clearly designed cognitive tasks, and behavioral outputs.
What is your favorite mentoring memory—either a story about a mentor’s impact on you or your impact on a mentee?
I can’t speak for my mentees in terms of my impact on them, and I leave them the freedom to have honest private thoughts. I myself had wonderful mentors throughout my career starting from the freshman years. I can focus on two favorite mentor stories. I call them the “two Jims”. My PhD advisor, Jim #1, James Byrne, (unfortunately deceased in 2022 before I could visit him one last time), was an amazing all-round renaissance man. His experience on the beamline was unbeatable, despite his scholarly appearance. But he also had superb theory knowledge and was a cultured and erudite man (although he had a famous Irish temper). He was the best writer I had ever encountered and could edit any of my clumsy scribblings down to half their original length and somehow make it sound elegant. His textbook about neutrons is still a classic in the field of low energy particle physics. Jim #2, who I enjoyed working with and discussing every conceivable issue with to this day, is James R. Moeller, the pioneer of the SSM-PCA technique, who was one of my mentors when I started my Postdoc at Columbia University many years ago. Jim #2 had an amazing and uncanny intuition that I sometimes struggled to understand and follow, but I brought Physics-inspired Monte-Carlo simulations to the table which helped test our ideas about high dimensional Euclidean spaces. I always enjoyed playing with simulated data and testing ideas – something I find useful to this day. In this manner of brainstorming and hashing out ideas and testing them in Monte Carlo simulations and applications to real data, Jim and I worked very fruitfully together with our complementary skill sets.
What scientist or scientific achievement do you most admire?
The overall achievement I would give the most credit to humanity’s well-being is the enlightenment and all the figures who contributed and are too numerous to mention. As for people, let me avoid the well-trodden references to Newton, Darwin, and Einstein – and pick some others who are also deserving. Let’s start with Alexander von Humboldt, a great 19th century explorer and polyglot (who inspired countless people, Goethe, Thomas Jefferson, Charles Darwin, Ernst Haeckel, and Simon Bolivar among them) and founded the field of wildlife ecology. Second, let’s pick Freeman Dyson as a 20th century genius. His accomplishments in showing the equivalency of three different formulations of Quantum Electrodynamics did not earn him the Nobel Prize, but he also had countless other interests and contributions. His essays are a delight to read. He was a Humboldt in the 20th Century.
Molecular connectivity: Best practices for data analysis
Bordeaux June 19th, 2026 – (In-person or virtually)
Registration: Register Here! Free of Charge!
Program & Speakers:
8:30 – 08:40 Welcome & Introduction by the organizers
08:40 – 09:10 (30 min) Molecular connectivity in the broader context of fMRI and other modalities –Bratislav Misic, McGill University (Canada)
09:10 – 9:30 (20 min) Introduction to molecular connectivity and nomenclature in the context of the Delphi study – Sharna Jamadar, Monash University (Australia)
9:30 – 9:50 (20 min) Overview of commonly used methods for assessment of molecular connectivity with emphasis on technical aspects that require discussion – Mattia Veronese, University of Padua (Italy)
09:50 – 10:10 (20 min) Preprocessing: Data harmonization, PVC, normalization – Martin Norgaard, University of Copenhagen (Denmark)
10:10 – 10:30 Coffee break
10:30 – 10:50 (20 min) General prerequisites (population heterogeneity, statistical power), minimum number of subjects for inter- and intra-subject estimation – Arianna Sala, Université de Liège (Belgium)
10:50 – 11:10 (20 min) ROI-level estimation metrics: partial or Pearson correlation, Euclidean Similarity – Tommaso Volpi, Yale University (USA)
11:10 – 11:30 (20 min) Voxel-level estimation: SSM-PCA vs. ICA (which method and when? selection of components), seed-based correlation – Matthieu Doyen, Université de Lorraine (France)
11:30 – 11:50 (20 min) Best practices for merging molecular, functional information and clinical info – Vince Calhoun, GSU, GATech, Emory University (USA)
11:50 – 12:10 (20 min) Statistical robustness (bootstrapping, corrections) – Chris Habeck, Columbia University (USA)
12:10 – 13:00 Summary and plan for future steps

📝Network-based disease fingerprinting with neuroinflammation PET imaging
The goal of this study by Barzon and colleagues was to develop a data-driven, network-based approach to generate individual brain-wide TSPO PET matrices, employing Euclidean distance to quantify inter-regional pharmacokinetics similarity. They applied this approach to a large multicenter dataset of 528 PET scans utilizing three different TSPO tracers, including healthy controls and patients with different diseases.
Read the full study in Journal of Neuroinflammation.
Key Findings:
📝Personalized mapping of body homeostasis using whole-body PET connectomics and routine FDG PET imaging
Labarthe and colleagues introduced a framework for generating individualized PET-based connectomes, enabling robust assessment of personalized systemic homeostasis.
Read the full study in Communications Medicine.
Key Findings:
📝Metabolic brain connectivity analysis of a depressive-like phenotype in rats: a graph theory PET study
In this study, Vazquez-Matias and colleagues aimed to investigate whether there are metabolic connectivity alterations in the brain of rats with a depressive-like phenotype, using positron emission tomography (PET) and graph theory methods. Male Wistar rats were exposed to 5 days of repeated social defeat (RSD) to induce a depressive-like phenotype, and brain connectivity was assessed with [18F]FDG-PET.
Read the full study in Psychiatry Res.
Key Findings:
📝Neuro–visceral immunometabolic network phenotyping via baseline whole-body [¹⁸F]FDG PET/CT underlying pathologic response to neoadjuvant immunochemotherapy in resectable NSCLC
In this study by Shi and colleagues, whole-body FDG-PET was used to compare cerebral metabolic networks between non-small cell lung cancer patients with pathological response to immunotherapy and patients that did not respond, as well as the characterisation of the brain-visceral metabolic coupling. Brain regions were grouped into two pre-defined cerebral modules: the Warburg-related module (regions with consistently high physiological glucose metabolism), and the anxiety/stress-related module.
Read the full study in European Journal of Nuclear Medicine and Molecular Imaging.
Key Findings:
📝[18F]Fluorodeoxyglucose positron emission tomography ([18F]FDG PET) characterises neurodegeneration levels across the -synucleinopathy continuum
Orso and colleagues aimed at assessing the ability of previously described brain metabolic disease-related covariance patterns (RP) to characterise neurodegeneration across the alpha-synuclein continuum, as well as in the specific clinical phenotypes (i.e., cognitive- and motor-predominant). To this end, they included healthy controls and isolated rapid eye movement sleep behaviour disorder, Parkinson’s disease (PD), and dementia with Lewy bodies patients.
Read the full study in Movement Disorders.
Key Findings:
📝Mapping individual molecular connectomes in Alzheimer’s disease
In this study, Xu and colleagues characterised the individual variations in amyloid tau pathology across the Alzheimer’s disease continuum by building individual molecular connectomes that capture the unique variations in pathological deposits in every patient. This new proposed measure of connectome alteration shows high discriminative ability both at baseline and in longitudinal analyses and correlates with cognitive decline over time more strongly than conventional SUVR metrics.
Read the full study in Alzheimer’s and Dementia.
Key Findings:
The MCWG Outreach Council invites you to submit announcements or information about papers, conferences, presentations or other events or news related to brain and molecular connectivity as well as any positions available or job opportunities that you wish to publicize and share with the community!
Please submit any material for consideration by the final day of each month using this form – thank you!

The MCWG is made up of four international and multidisciplinary councils dedicated to promoting molecular connectivity research via dissemination of methods, results, collaboration, and resource sharing (e.g. datasets, tools) within the scientific community. We encourage the neuroscientific community to take an integrative perspective in study of the brain connectome, where various methods including MRI-based techniques, electrophysiological tools, and molecular imaging advance our understanding of the brain. Please find fundamental questions outlined here: “Brain connectomics: time for a molecular imaging perspective?”
Our website can be found here. We also invite you to join the MCWG!
[MCWG] Molecular Connectivity Newsletter: April 2026

Thank you to everyone who joined us for March’s MCOS with Penghui Du! Missed it? The recording is available here.
Wishing our community a joyful Easter filled with renewal, inspiration, and continued progress in advancing brain molecular connectivity!

Date: April 17th, 2026
Time: 13:00 UTC
Registration: Please register here.
Title: Preoperative network activity predicts the response to subthalamic DBS for Parkinson’s disease
Speaker: Dr. Prashin Unadkat
Abstract: Does the brain’s baseline network architecture tell us about an individual’s capacity to respond to neuromodulation? Can we use this to identify which Parkinson’s disease patients will benefit most from deep brain stimulation before they ever reach the operating room? This talk will present our work developing and validating STN StimNet, a treatment-specific metabolic brain network identified in PD patients with implanted subthalamic nucleus (STN) electrodes. Several key questions will be addressed: What distinguishes a treatment-induced network from a disease-related one? What is the electrophysiological basis of this network; specifically, why do STN theta-band oscillations, rather than the pathological beta activity traditionally linked to PD motor impairment, drive StimNet expression? We go on to demonstrate how preoperatively measured network expression with either FDG PET or resting-state fMRI predicts postoperative motor outcomes, and how it compares to the current gold standards for patient selection. Using data from a large cohort of PD patients spanning the clinical spectrum of the disease, we illustrate how these measurements can stratify patients by their likelihood of achieving a clinically meaningful response, while appropriately excluding populations unlikely to benefit, such as those with atypical parkinsonian syndromes. Finally, this predictive framework may have relevance beyond STN-DBS in Parkinson’s disease. The principle that preoperative circuit-level activity shapes an individual’s response to focal neuromodulation could guide patient selection across other DBS targets and may extend to other movement and psychiatric disorders.

Dr. Prashin Unadkat, MBBS, PhD is the Chief Resident in the Department of Neurosurgery at the Zucker School of Medicine at Hofstra/Northwell and an incoming Stereotactic and Functional Neurosurgery fellow at Baylor College of Medicine. His clinical practice focuses on image-guided and neuromodulatory surgical strategies for movement disorders, psychiatric conditions, and epilepsy. His research centers on developing functional and structural brain imaging biomarkers that can predict treatment response and improve surgical planning, with a particular emphasis on network-level analysis of deep brain stimulation effects.
The MCOS promotes rigor in research and resource sharing. We aim to hold MCOS every third Friday of the month, subject to change due to speaker availability. Please stay tuned for MCOS updates and reminders on social media! Thank you!
Each month, we will feature a member of the MCWG and have a brief Q&A!
This month please enjoy our highlight of Prof Dr Daniela Perani, member of the MCWG Steering Committee.

Prof Dr Daniela Perani is an internationally renowned neurologist and neuroscientist, and Professor Emeritus of Neuroscience at the Università Vita-Salute San Raffaele in Milan. She has served as the coordinator of numerous national and international research projects. Her work in the field of cognitive neuroscience has extensively employed neuroimaging techniques to investigate language, memory, and the perception of music. In the clinical domain, she has focused in particular on the neural correlates of neurodegenerative diseases. She has authored more than 400 peer-reviewed publications in international journals, as well as numerous book chapters and books.
Prof Dr Daniela Perani has graciously responded to our feature questionnaire:
What sparked your interest in molecular imaging or led you to focus on research in molecular imaging?
My interest in molecular imaging was sparked early in my career through my work in neuroscience, particularly in the study of brain function and cognitive processes in health and diseases. During my initial training and research activities, I became deeply interested in understanding how complex neurological and cognitive functions—such as language, memory, and neurodegenerative changes—could be linked to underlying biological mechanisms in the living human brain. At that time in the ninety, molecular imaging represented a unique and emerging opportunity to bridge neuroscience and clinical research. My work focused on applying techniques such as MRI and PET to investigate brain anatomy, metabolism and neurotransmitter systems, allowing me to explore both neurological disorders and cognitive processes in vivo. This approach enabled me to connect molecular and functional alterations with clinical symptoms, especially in conditions like dementia and language disorders. What motivated me most was the possibility of combining advanced imaging technologies with questions about how the brain supports cognition, and how these processes are disrupted in disease. This intersection between molecular mechanisms and neurological and cognitive function has remained a central theme throughout my career.
What is your role in the Molecular Connectivity Working Group, and what have you been contributing to or working on within the group?
Within the Molecular Connectivity Working Group, I currently serve as a member of the Board of Directors, contributing in a senior and advisory capacity. My involvement builds on my earlier work, where I conducted some of the pioneering studies on PET-based molecular connectivity, helping to establish the conceptual and methodological framework that has guided much of the subsequent research in this area. These initial contributions were aimed at demonstrating how molecular imaging, and in particular PET, could be used not only to assess regional brain activity but also to investigate connectivity patterns at a molecular level. This work contributed to opening new directions for understanding brain organization and dysfunction.
In my current role, I remain actively engaged in the group’s activities by providing critical input on ongoing projects, contributing to strategic discussions, and offering guidance on methodological developments. I also support the group through active suggestions and mentorship, helping to ensure scientific rigor and to foster the continued evolution of molecular connectivity research.
In what ways do you imagine molecular connectivity will advance our understanding of brain function?
Molecular connectivity could greatly enhance our understanding of brain function by linking large-scale brain networks to their underlying biochemical processes. Unlike MRI, which captures structural or functional activity indirectly, PET provides insight into metabolism and neurotransmitter systems, bringing us closer to the brain’s biological foundations. However, challenges remain. For example, FDG-PET connectivity is often based on correlations in glucose uptake, whose biological meaning is still unclear — they may reflect true neural interactions, shared inputs, or global effects. Differences between PET and MRI networks further highlight the need for careful multimodal integration. Methodological issues also persist, including sensitivity to noise, variability, and preprocessing choices. These require more robust statistical approaches. The complexity increases with neurotransmitter-based connectivity, where correlations in receptor binding do not necessarily indicate direct functional links. Overall, the key goal is to connect molecular connectivity findings to clear biological interpretations. If these challenges are addressed, it could bridge molecular and systems neuroscience, offering deeper insight into cognition and brain disorders.
What do you think are the most important challenges in current brain connectivity research, or which unsolved/underappreciated issues should the community address?
Brain connectivity research faces several key challenges that must be addressed to achieve true biological insight. A central issue is that connectivity is still mostly inferred from statistical relationships (e.g., correlations) rather than direct neural interactions, making interpretation uncertain—especially in fMRI and PET. Another major challenge is integrating different imaging modalities. MRI and PET capture distinct aspects of brain function (hemodynamic vs. biochemical), but their results often only partially overlap, highlighting the lack of a unified framework. Similarly, molecular and neurotransmitter connectivity can be difficult to interpret, as correlations in receptor binding do not necessarily reflect functional interactions. Neurophysiology (e.g., EEG/ERP) offers an important opportunity by providing fast temporal dynamics that can help link and validate findings across modalities, potentially enabling more mechanistic models of brain function. Methodological limitations also persist, including sensitivity to noise, variability across subjects and sites, and reliance on simplified models. There is a strong need for large, standardized, multimodal datasets. While AI can help analyze complex data, its success depends on high-quality data and biologically meaningful questions—otherwise, it risks producing results that are technically advanced but hard to interpret.
What is your favorite mentoring memory—either a story about a mentor’s impact on you or your impact on a mentee?
I don’t have a “favorite mentoring memory”. In the fields like brain connectivity and molecular neuroscience, a powerful mentoring memory is when a mentor helps someone move from feeling overwhelmed by complexity to seeing a clear path forward. For example, a student working on brain connectivity at the molecular level might be struggling to integrate vast datasets—gene expression, synaptic markers, and network-level imaging. A good mentor doesn’t just give answers; they help the mentee reframe the problem, connect concepts across scales, and build confidence in handling ambiguity.
What scientist or scientific achievement do you most admire?
First, I want to underline that I admire joy, enthusiasm, and tenacity, as well as the ability to face doubts, support young people in their research, and share results without ever overpowering others. I admire all the work carried out by Eric Kandel all over his scientific career and his group.
Molecular connectivity: Best practices for data analysis
Bordeaux June 19th, 2026 – (In-person or virtually)
Registration: Register Here! Free of Charge!
Program & Speakers:
8:30 – 08:40 Welcome & Introduction by the organizers
08:40 – 09:10 (30 min) Molecular connectivity in the broader context of fMRI and other modalities –Bratislav Misic, McGill University (Canada)
09:10 – 9:30 (20 min) Introduction to molecular connectivity and nomenclature in the context of the Delphi study – Sharna Jamadar, Monash University (Australia)
9:30 – 9:50 (20 min) Overview of commonly used methods for assessment of molecular connectivity with emphasis on technical aspects that require discussion – Mattia Veronese, University of Padua (Italy)
09:50 – 10:10 (20 min) Preprocessing: Data harmonization, PVC, normalization – Martin Norgaard, University of Copenhagen (Denmark)
10:10 – 10:30 Coffee break
10:30 – 10:50 (20 min) General prerequisites (population heterogeneity, statistical power), minimum number of subjects for inter- and intra-subject estimation – Arianna Sala, Université de Liège (Belgium)
10:50 – 11:10 (20 min) ROI-level estimation metrics: partial or Pearson correlation, Euclidean Similarity – Tommaso Volpi, Yale University (USA)
11:10 – 11:30 (20 min) Voxel-level estimation: SSM-PCA vs. ICA (which method and when? selection of components), seed-based correlation – Matthieu Doyen, Université de Lorraine (France)
11:30 – 11:50 (20 min) Best practices for merging molecular, functional information and clinical info – Vince Calhoun, GSU, GATech, Emory University (USA)
11:50 – 12:10 (20 min) Statistical robustness (bootstrapping, corrections) – Chris Habeck, Columbia University (USA)
12:10 – 13:00 Summary and plan for future steps

👨🏼🔬 We are looking for Volunteers – Join us!
The resources committee is currently looking for volunteers for the literature review of studies in molecular connectivity.
If you are interested in joining us, please reach out through: https://molecularconnectivity.com/how-to-join/
📝Task-evoked brain network architecture captured by the complementary integration of metabolic and functional connectivity
The goal of this study by Vallini and colleagues was to map the reconfiguration of the whole brain metabolic network in response to increasingly difficult tasks, and to investigate how both functional and metabolic reconfigurations contribute to behavioural performance during the task.
Read the full study in NeuroImage.
Key Findings:
📝Individual-level metabolic connectivity captures cortical morphology and their coupling strengthens with age
In this study by Facca and colleagues, the integration of FDG PET and structural MRI data were used to investigate how cortical morphology constrains large-scale patterns of metabolic connectivity across the human lifespan.
Read the full study in bioRxiv.
Key Findings:
📝Utility of [18F]PI-2620 as universal biomarker for the amyloid/tau/neurodegeneration classification of Alzheimer disease: an exploratory study with dual-phase PET imaging
Franceschi and colleagues aimed at evaluating the efficacy of [18F]PI-2620 as a single biomarker for the A/T/N classification through the identification of spatial covariance patterns using SSM/PCA in early- and late-phase scans of amyloid-positive Alzheimer’s disease patients and amyloid-negative healthy controls.
Read the full study in Neuroradiology.
📝 Novel insights into cognitive network alterations in temporal lobe epilepsy: a [18F]SynVesT-1 PET study
Qui and colleagues employed the Kullback-Leibler Divergence Similarity Estimation algorithm to construct individualised brain synaptic connectomes using [18F]SynVesT-1 PET imaging in temporal lobe epilepsy patients. Their main hypothesis was that distributed synaptic connectivity attenuation, rather than focal synaptic loss, mediates cognitive network dysfunction in epilepsy.
Read the full study in Epilepsy & Behaviour.
The MCWG Outreach Council invites you to submit announcements or information about papers, conferences, presentations or other events or news related to brain and molecular connectivity as well as any positions available or job opportunities that you wish to publicize and share with the community!
Please submit any material for consideration by the final day of each month using this form – thank you!

The MCWG is made up of four international and multidisciplinary councils dedicated to promoting molecular connectivity research via dissemination of methods, results, collaboration, and resource sharing (e.g. datasets, tools) within the scientific community. We encourage the neuroscientific community to take an integrative perspective in study of the brain connectome, where various methods including MRI-based techniques, electrophysiological tools, and molecular imaging advance our understanding of the brain. Please find fundamental questions outlined here: “Brain connectomics: time for a molecular imaging perspective?”
Our website can be found here. We also invite you to join the MCWG!
[MCWG] Molecular Connectivity Newsletter: March 2026

Thank you to everyone who joined us for February’s MCOS with Dr. Paul Klauser and Dr. Federico Lucchetti! Missed it? The recording is available here.
Date: March 20th, 2026
Time: 14:00 UTC
Registration: Please register here.
Title: Human Cerebral Cortex Organization Characterized by Functional PET-FDG “Metabolic Connectivity”
Speaker: Penghui Du
Abstract: Resting-state metabolic connectivity (RSMC) as measured by [18F]-fluorodeoxyglucose functional PET (fPET-FDG) provides a promising framework for understanding the brain’s energetic organization, yet its spatiotemporal structure remains insufficiently characterized. Here, we investigated the cortical organizational principles of RSMC in the human brain and their relationship to other known cortical organizational features. At the local scale, metabolic boundaries partly overlapped with resting-state functional connectivity patterns. At the global scale, RSMC was organized along a robust superior-inferior cortical gradient, and was primarily driven by low-frequency, minute-scale fPET-FDG dynamics. This large-scale metabolic profile aligned with known anatomical and energetic constraints, providing new insights into the brain’s energetic architecture. Link to BioRxiv

Penghui Du is currently a Master’s student in Neuro-X at EPFL. He received his BSc in Intelligent Medical Engineering from the Southern University of Science and Technology in 2024. As a visiting student at the CANDY Lab (Athinoula A. Martinos Center for Biomedical Imaging, Massachusetts General Brigham, United States), he worked with Dr. Jingyuan Chen on mapping the organization of the human cerebral cortex using functional PET-FDG metabolic connectivity. Personal website: https://penghui-du.com/
The MCOS promotes rigor in research and resource sharing. We aim to hold MCOS every third Friday of the month, subject to change due to speaker availability. Please stay tuned for MCOS updates and reminders on social media! Thank you!
Each month, we will feature a member of the MCWG and have a brief Q&A!
This month please enjoy our highlight of Prof Dr Vesna Sossi, member of the MCWG Steering Committee.

Prof Dr Vesna Sossi received a Laurea degree in high energy Physics from the University of Trieste Italy and a PhD in Nuclear Physics from the University of British Columbia in 1991. After completing her graduate degree, she went on to a post-doctoral fellowship in the UBC/TRIUMF PET group working on Medical Imaging. She is currently a Professor in the Physics and Astronomy Department and Adjunct Professor in Medicine at UBC and has been leading the UBC Positron Emission Tomography (PET), now PET/MRI brain imaging program since 2009.
Professor Sossi’s expertise and research interests lie in using clinical and preclinical imaging to investigate neurodegeneration and other brain diseases through development of instrumentation, data quantification, image analysis, kinetic modelling, image reconstruction and novel imaging protocols. She is particularly interested to further develop and exploit hybrid PET/MR imaging to gain access to as yet poorly investigated aspects of brain function such as brain energetics and neurovascular coupling in the healthy brain and as affected by neurodegeneration and exercise, as well as other possible neuroprotective mechanisms. Some examples of her research are: the development of a complex dopamine turnover measurement using PET in humans and rodents and the demonstration of its relevance to Parkinson’s disease (PD), first applications of texture and pattern analyses to PET imaging of several neurotransmitter systems, particularly well-suited to identify interactions between systems, and lately exploring brain energetics and its modulation by exercise in PD. She has been elevated to IEEE Fellow status for ‘for contributions to quantitative and translational brain PET imaging’.
She has over 250 peer reviewed journal publications, and has trained many graduate and undergraduate students. Professor Sossi sits on several national and international review committees and received several CFI, NSERC and Michael Smith Foundation for Health Research awards during her career. She is also currently Director Elect for IEEE Division IV.
Prof Dr Vesna Sossi has graciously responded to our feature questionnaire:
What sparked your interest in molecular imaging or led you to focus on research in molecular imaging?
I have originally chosen physics as my field of studies, as I was fascinated by the world around me. However I have been equally interested in medicine and particularly in how the brain works. Molecular imaging provided a natural bridge between the two through radiation detectors. The more I learned about the power of molecular imaging, the more I was intrigued by the unique information it could provide on brain function.
In what ways do you imagine molecular connectivity will advance our understanding of brain Function?
Molecular connectivity promotes thinking of the brain as a very complex circuit with feedback loops and interconnected effects, which, I believe, better represents the intricate functioning of the brain than looking at it as a composite of individual entities. Indeed, many disease and intervention related effects can be more readily observed on a pattern level rather than investigating behaviours of individual brain regions separately. Connectivity type analysis also lends itself well to inclusion of multi-modal data, possibly revealing related effects in different brain function and neurochemical domains.
What do you think are the most important challenges in current brain connectivity research, or which unsolved/underappreciated issues should the community address?
Lack of uniform imaging protocols including choice of radiotracers, data acquisition and processing, and possibly subject selection criteria. Equally important is validation of the methodology, analysis and statistical approaches, pipelines and cutoff criteria, when relevant. Another challenge is how to best relate the effects observed at a spatial and temporal scale commensurate with imaging to the underlying effects occurring at the cellular level which operate at different scales and generally involve a much larger degree of complexity.
What is your favorite mentoring memory—either a story about a mentor’s impact on you or your impact on a mentee?
There are many stories, but one of my favourite ones dates to the very early years of my molecular imaging career, when I was writing my first paper in this area. I wrote ‘… I placed a region of interest on the striatum….’. Dr. Brian Pate, who was at that time leading the UBC Primate imaging program, looked at me and said: ‘No, this is not what you have done’. He must have registered the confused look on my face and said with a chuckle: ‘ You placed a region of interest on the IMAGE of the striatum!’. It may appear insignificant, but that comment prompted me to examine extremely carefully what is being done at each step of any analysis approach for the rest of my career – and not just when it came to descriptions!
Multimodal Integration in Human Brain Mapping
Bordeaux June 14, 2026 – (In-person at Palais 2 l’Atlantique – duration: 4h)
Organizers: Prof Dr Joana B Pereira and Dr Arianna Sala
Capturing Rich Multimodal Brain Interactions: Model Selection, Interpretability, and Clinical Applications (20 min)
Prof. Jing Sui – IDG/McGovern Institute for brain research, State Key Laboratory of Cognitive Neuroscience and Learning, Beijing Normal University, China
Neuromaps: A Python Toolbox for Cross-Modal Standardization and Interpretation (20 min)
Dr. Vincent Bazinet – Montréal Neurological Institute, McGill University, Montréal, Quebec, Canada
Linking MRI-Derived Measures and Underlying Neurophysiology Using the JuSpace Toolbox (20 min)
Prof. Juergen Dukart – Institute of Neurosciences and Medicine, Brain & Behaviour (INM-7), Research Centre Juelich; Juelich, Germany
Multilayer Network Modelling with the BRAPH 2 software (20 min)
Assoc. Prof. Joana B. Pereira – Clinical Neurosciences Department, Kalinska Institute, Stockholm, Sweden
Integrating fMRI and PET data through REACT (20 min)
Dr. Manuela Moretto – Department of Information Engineering, University of Padua, Padua, Italy
JuSpace: Spatial Correlation Analyses for Molecular–MRI Integration (35 min)
Prof. Juergen Dukart – Institute of Neurosciences and Medicine, Brain & Behaviour (INM-7), Research Centre Juelich; Juelich, Germany
Practical Tutorial with Neuromaps (35 min)
Dr. Vincent Bazinet – Montréal Neurological Institute, McGill University, Montréal, Quebec, Canada
Multilayer Network Analyses with BRAPH 2 (35 min)
MS Yu-Wei Chang – Department of Physics, University of Gothenburg, Gothenburg, Sweden
Receptor-Enriched Functional Connectivity with REACT (35 min)
Dr. Manuela Moretto – Department of Information Engineering, University of Padua, Padua, Italy
👨🏼🔬 We are looking for Volunteers – Join us!
The resources committee is currently looking for volunteers for the literature review of studies in molecular connectivity.
If you are interested in joining us, please reach out through: https://molecularconnectivity.com/how-to-join/
📝A Data-Driven SSM/PCA Analysis Approach for Differential Diagnosis of Parkinsonism Using 11C-PE2I PET
This study by Falk and colleagues was a proof-of concept study to derive specific disease patterns from parametric images derived from dynamic 11C-PE2I (dopamine transporter imaging) images for differential diagnosis of individuals with Parkinson’s disease, dementia with Lewy bodies, and progressive supranuclear palsy.
Read the full study in NeuroImage Clinical.
Key Findings:
📝Altered brain glucose metabolism and connectivity in young adults with obstructive sleep apnea
In this study by Caminiti and colleagues the effects of moderate to severe obstructive sleep apnea syndrome on the brain were analysed using FDG PET comparing patients to cognitively unimpaired or without other systemic or neurological disorders individuals. A voxel-wise seed-based interregional correlation analysis was performed to assess large-scale networks affected by the disease.
Read the full study in Alzheimer’s and Dementia.
Key Findings:
📝Early diagnosis of Alzheimer’s disease: graph theoretical analysis of cerebellar network features based on [18F]AV45 PET
Li and colleagues had the primary objective to investigate changes in cerebellar amyloid plaques along the Alzheimer’s disease spectrum. Using amyloid PET and graph theory, they concluded that abnormal amyloid plaque deposition affects connectivity networks in the early stages of Alzheimer’s disease.
Read the full study in PLoS One.
📝Human Cerebral Cortex Organization Characterized by Functional PET-FDG “Metabolic Connectivity”
In this study, Du and colleagues characterized the spatiotemporal organization of resting-state metabolic connectivity (RSMC) in the human brain, as measured by [18F]fluorodeoxyglucose (FDG) functional PET (fPET-FDG). They examined the relationship between RSMC organization and resting-state functional connectivity (RSFC) derived from functional magnetic resonance imaging and other known cortical organizational principles.
Read the full study in bioRxiv.
📝Network-Based Analysis for the Quantification of Brain and Body Immune Axes with Total-Body PET Imaging
In this study, Maccioni and colleagues aimed to validate network analysis of total-body 18-kDa translocator protein (TSPO) PET imaging for studying brain and body immune axes.
Read the full study in Journal of Nuclear Medicine.
Key Findings:
📝Enhancing diagnosis of mild cognitive impairment through brain-heart-gut metabolic networks in whole-body PET imaging
In this study, Li and colleagues presented a framework integrating brain-heart-gut interactions using whole-body positron emission tomography (PET) to enhance brain-only diagnostic performance in mild cognitive impairment (MCI).
Read the full study in Cell Reports Medicine.
Key Findings:
The MCWG Outreach Council invites you to submit announcements or information about papers, conferences, presentations or other events or news related to brain and molecular connectivity as well as any positions available or job opportunities that you wish to publicize and share with the community!
Please submit any material for consideration by the final day of each month using this form – thank you!

The MCWG is made up of four international and multidisciplinary councils dedicated to promoting molecular connectivity research via dissemination of methods, results, collaboration, and resource sharing (e.g. datasets, tools) within the scientific community. We encourage the neuroscientific community to take an integrative perspective in study of the brain connectome, where various methods including MRI-based techniques, electrophysiological tools, and molecular imaging advance our understanding of the brain. Please find fundamental questions outlined here: “Brain connectomics: time for a molecular imaging perspective?”
Our website can be found here. We also invite you to join the MCWG!
[MCWG] Molecular Connectivity Newsletter: February 2026
Thank you to everyone who joined us for January’s MCOS with Dr. Sarah Genon! Missed it? The recording is available here.
[MCWG] Molecular Connectivity Newsletter: January 2026

Thank you to everyone who joined us for November’s Special Symposium edition! Missed it? The recording is available here.
Date: January 23rd, 2026
Time: 14:00 UTC
Registration: Please register here.
Title: Connectivity-based parcellation to map brain organization
Speaker: Dr. Sarah Genon
Abstract: The human brain is often described in terms of discrete regions, yet defining brain atlases remains a central challenge in neuroscience. Connectivity-based parcellation offers a principled framework for identifying functionally coherent regions using a variety of connectivity markers. In this talk, Dr. Sarah Genon will highlight how metabolic connectivity can be leveraged to derive region definitions grounded in metabolic network organization. I will discuss the relevance of these connectivity-based regions for improving our understanding of brain–behavior relationships and characterizing dysfunction in clinical populations.

Dr. Sarah Genon is a cognitive neuroscientist specialized in neuroimaging, machine learning, and the study of brain–behavior relationships. She is a Heisenberg Professor at the Heinrich-Heine University of Dusseldorf and a group leader at the Forschungszentrum Jülich (Germany).
The MCOS promotes rigor in research and resource sharing. We aim to hold MCOS every third Friday of the month, subject to change due to speaker availability. Please stay tuned for MCOS updates and reminders on social media! Thank you!
Each month, we will feature a member of the MCWG and have a brief Q&A!
This month please enjoy our highlight of Prof. Dr. Kristina Herfert, member of the MCWG Steering Committee.

Prof Dr Kristina Herfert is an Associate Professor for Functional and Metabolic Brain Imaging at Werner Siemens Imaging Center, Department of Preclincal Imaging and Radiopharmacy, University of Tübingen, Germany. Her research group’s aim is to develop and apply protocols and methods to assess molecular changes of receptor and protein expression by PET and functional changes by BOLD-fMRI to develop early read-outs of disease progression. Her group’s current research topics are: Hybrid Imaging Using PET and fMRI to Study Brain Functional Connectivity, Quantitative Functional and Molecular Brain Imaging in Animal Models of Neurodegenerative Disorders and PET Radiotracer Development in Brain Neurodegenerative Diseases.
Prof Dr. Kristina Herfert has graciously responded to our feature questionnaire:
What sparked your interest in molecular imaging or led you to focus on research in molecular imaging?
I have always been deeply curious about how the brain works and how it is able to perform such extraordinarily sophisticated functions. As the most complex organ in the human body, the brain has fascinated me for as long as I can remember. While in vitro studies using cell models and tissue have been instrumental in advancing our understanding of molecular disease mechanisms, I have always been particularly drawn to studying the brain in vivo, in its natural, functioning state. Only under in vivo conditions can we truly capture the dynamic interplay between different cell types, receptors, metabolism, and signaling pathways. PET imaging uniquely enables us to investigate these processes noninvasively using a wide range of molecular tracers. The continuous development of new tracers further fuels my interest, as it constantly opens up new possibilities to study brain function and neurological disease at the molecular level.
What is your role in the Molecular Connectivity Working Group, and what have you been contributing to or working on within the group?
I became a member of the Molecular Connectivity Working Group, because I am highly motivated to explore molecular connectivity as a way to study the brain and its neurological disorders beyond conventional PET quantification approaches. A key contribution I hope to bring to the group is my background in preclinical imaging, which allows for the integration of molecular connectivity with methods that offer higher specificity and resolution. In particular, combining molecular connectivity analyses with genetic approaches, such as genetically encoded neurotransmitter sensors, optogenetic manipulation or animal models, can provide deeper insights into specific pathways and molecular mechanisms with superior temporal and spatial resolution. By linking molecular connectivity measures with more invasive, mechanistic readouts, I see strong potential for a multimodal imaging framework that can help validate and better interpret molecular connectivity findings. Within the working group, I am eager to exchange ideas, collaborate internationally, and help advance this integrative perspective.
In what ways do you imagine molecular connectivity will advance our understanding of brain function?
Brain connectivity has fundamentally changed how we think about brain organization by shifting the focus from isolated brain regions to distributed networks. Molecular connectivity has the potential to bring this network perspective to the molecular level. PET imaging enables the investigation of a wide range of processes, including metabolism, neuroinflammation, receptor availability, and neurotransmitter dynamics. Although PET has lower spatial and temporal resolution compared to fMRI, its molecular specificity and sensitivity are unique strengths. Molecular connectivity therefore offers the opportunity to study how molecular processes are coordinated across brain networks, rather than locally confined. While further methodological development is needed to ensure that these network-level molecular measures are robust and biologically meaningful, I strongly believe that molecular connectivity will substantially deepen our understanding of brain function in both health and disease.
What do you think are the most important challenges in current brain connectivity research, or which unsolved/underappreciated issues should the community address?
One of the central challenges in functional connectivity research is that the BOLD signal is an indirect, hemodynamic measure rather than a direct reflection of neuronal activity, making it sensitive to vascular confounds. In addition, connectivity analyses often attempt to relate macroscale network measures to microscale biological processes such as synaptic function, receptor expression, or glial activity. As a result, changes in connectivity can reflect multiple overlapping mechanisms, which complicates interpretation. PET-based molecular connectivity offers greater specificity to particular molecular processes, but is limited by relatively low temporal resolution, making it difficult to capture fast network dynamics. More fundamentally, we still do not fully understand what “connectivity” represents biologically. Molecular coupling may arise from shared receptor expression, similar cell-type composition, common vascular or metabolic constraints, or developmental gradients. Addressing these conceptual and methodological challenges will require integrative multimodal approaches and more preclinical studies that link molecular connectivity measures to known biological mechanisms.
What is your favorite mentoring memory—either a story about a mentor’s impact on you or your impact on a mentee?
One of the most formative mentoring relationships in my career has been with Prof. Bernd Pichler, the director of our institute. His enthusiasm for imaging and his deep curiosity about science have had a lasting influence on my development as a researcher. As a PhD student, I often entered meetings with him feeling discouraged, convinced that my results were not good enough or that my data were insufficient. Yet I almost always left those meetings feeling energized and confident. He has a rare ability to motivate people and to reframe challenges as opportunities for discovery. Equally important, he has always given me a great deal of freedom in pursuing my own research ideas, encouraging independence and creativity. This balance of support, trust, and intellectual freedom has strongly shaped how I think about research and mentorship today.
What scientist or scientific achievement do you most admire?
This is a difficult question, as most scientific advances today are the result of collaborative efforts rather than individual achievements. For that reason, it feels limiting to single out one contemporary scientist. However, when reflecting on the historical context of science, particularly the challenges faced by women, Marie Curie is someone I deeply admire. Her pioneering work laid the foundation for nuclear medicine, and she was the first woman to receive the Nobel Prize—remarkably in two different disciplines, physics and chemistry. Her scientific legacy and perseverance continue to make her a powerful role model for women in science.
Molecular Connectivity: Best practices for data analysis
Bordeaux June 19, 2026
8:30 – 08:40 Welcome & Introduction by the organizers
08:40 – 09:10 (30 min) Molecular connectivity in the broader context of fMRI and other modalities
Bratislav Misic, McGill University (Canada)
09:10 – 9:30 (20 min) Introduction to molecular connectivity and nomenclature in the context of the Delphi study
Sharna Jamadar, Monash University (Australia)
9:30 – 9:50 (20 min) Overview of commonly used methods for assessment of molecular connectivity with emphasis on technical aspects that require discussion
Mattia Veronese, University of Padua (Italy)
09:50 – 10:10 (20 min) Preprocessing: Data harmonization, PVC, normalization
Martin Norgaard, University of Copenhagen (Denmark)
10:10 – 10:30 Coffee break
10:30 – 10:50 (20 min) General prerequisites (population heterogeneity, statistical power), minimum number of subjects for inter- and intra-subject estimation
Arianna Sala, Université de Liège (Belgium)
10:50 – 11:10 (20 min) ROI-level estimation metrics: partial or Pearson correlation, Euclidean Similarity
Tommaso Volpi, Yale University (USA)
11:10 – 11:30 (20 min) Voxel-level estimation: SSM-PCA vs. ICA (which method and when? selection of components), seed-based correlation
Matthieu Doyen, Université de Lorraine (France)
11:30 – 11:50 (20 min) Best practices for merging molecular, functional information and clinical info
Vince Calhoun, GSU, GATech, Emory University (USA)
11:50 – 12:10 (20 min) Statistical robustness (bootstrapping, corrections)
Chris Habeck, Columbia University (USA)
12:10 – 13:00 Summary and plan for future steps
The [18F]FDG-PET Workshop
Assessing Brain Glucose Metabolism in Patients with Disorders of Consciousness
From Acquisition to Interpretation aims to provide a comprehensive overview of the use of [18F]FDG-PET imaging in brain-injured patients. Speakers will cover a wide range of topics, including patient preparation, tracer kinetics, data acquisition, image processing and analysis, and the interpretation of clinical results.
The workshop will conclude with a forward-looking presentation on the future of PET imaging and its implications for patients with disorders of consciousness. A major emphasis will be placed on hands-on activities to support practical learning, as well as on the presentation of real clinical cases, fostering discussion and interaction between participants and experts.
The faculty includes Dr. Jitka Annen (University of Ghent), Ir. Claire Bernard (University Hospital of Liège), Dr. Florentin Kucharczak (University of Montpellier), Dr. Arianna Sala (University of Liège), and Dr. Tommaso Volpi (Yale University).
The detailed information of the event and free registration are available here.
📝 Metabolic brain networks in dementia with Lewy bodies: from prodromal to manifest disease stages
In this study, Perovnik et al analysed a large multicenter FDG PET dataset of more the 1,180 participants from 14 tertiary centers with prodromal and manifest dementia with Lewy bodies (DLB), Alzheimer’s disease (AD), and normal controls to evaluate the clinical utility of a quantifiable metabolic network biomarker, termed DLB-related pattern (DLBRP).
Read the full study in Journal of Neurology, Neurosurgery & Psychiatry.
Key Findings:
📝 Relationships between local metabolic activity and distributed functional connectivity in major depressive disorder
In this study, Sun and colleagues evaluated the interactions between local activity and distributed connectivity to reveal novel pathobiological signatures of Major Depressive Disorder (MDD).
Read the full study in Translational Psychiatry.
Key Findings:
📝 Constructing the human brain metabolic connectome with MR spectroscopic imaging reveals cerebral biochemical organization
In this study, Lucchetti and colleagues using fast, high-resolution 3D whole-brain proton magnetic resonance spectroscopic imaging (1H-MRSI), derived a within-subject metabolic connectome in 51 healthy subjects, defined as pairwise correlations among five metabolites (tCr, tNAA, Glx, Ins, Cho) across gray-matter parcels.
Read the full study in Nature Communications.
The MCWG Outreach Council invites you to submit announcements or information about papers, conferences, presentations or other events or news related to brain and molecular connectivity as well as any positions available or job opportunities that you wish to publicize and share with the community!
Please submit any material for consideration by the final day of each month using this form – thank you!

The MCWG is made up of four international and multidisciplinary councils dedicated to promoting molecular connectivity research via dissemination of methods, results, collaboration, and resource sharing (e.g. datasets, tools) within the scientific community. We encourage the neuroscientific community to take an integrative perspective in study of the brain connectome, where various methods including MRI-based techniques, electrophysiological tools, and molecular imaging advance our understanding of the brain. Please find fundamental questions outlined here: “Brain connectomics: time for a molecular imaging perspective?”
Our website can be found here. We also invite you to join the MCWG!
[MCWG] Molecular Connectivity Newsletter: December 2025

Thank you to everyone who joined us for November’s Special Symposium edition! Missed it? The recording is available here.
We would like to wish everyone a very happy holiday season and we send our wishes for a happy and healthy new year!

Date: January 23rd, 2026
Time: 14:00 UTC
Registration: Please register here.
Title: Connectivity-based parcellation to map brain organization
Speaker: Dr. Sarah Genon
Abstract: The human brain is often described in terms of discrete regions, yet defining brain atlases remains a central challenge in neuroscience. Connectivity-based parcellation offers a principled framework for identifying functionally coherent regions using a variety of connectivity markers. In this talk, Dr. Sarah Genon will highlight how metabolic connectivity can be leveraged to derive region definitions grounded in metabolic network organization. I will discuss the relevance of these connectivity-based regions for improving our understanding of brain–behavior relationships and characterizing dysfunction in clinical populations.

Dr. Sarah Genon is a cognitive neuroscientist specialized in neuroimaging, machine learning, and the study of brain–behavior relationships. She is a Heisenberg Professor at the Heinrich-Heine University of Dusseldorf and a group leader at the Forschungszentrum Jülich (Germany).
The MCOS promotes rigor in research and resource sharing. We aim to hold MCOS every third Friday of the month, subject to change due to speaker availability. Please stay tuned for MCOS updates and reminders on social media! Thank you!
📝 Functional PET for mapping metabolic dynamics in Parkinson’s disease
In this study, Heinecke and colleagues combined resting state functional 18F-FDG PET (fPET) and functional magnetic resonance imaging to evaluate differences between healthy controls and Parkinson’s disease (PD) patients in glucose consumption, establish a measure of glucose dynamics at the subject-level, and develop a seed-based network approach for analysing metabolic time series data.
Read the full study in Scientific Reports.
Key Findings:
📝 Fully Individualized Models for Cross-Sectional and Longitudinal Network-Based Tau Spread
[18F]Flortaucipir PET and diffusion magnetic resonance imaging were used to define individualized epicentres of tau pathology. Using data from the Alzheimer’s Disease Neuroimaging Initiative, Brown and colleagues showed that a model using individualised tau epicentres outperform models to assess tau covariance that are based on group epicentres and/or group connectomes.
Read the full study in Imaging Neuroscience.
Key Findings:
📝 Whole-Body [18F]FDG-PET/CT Imaging of Healthy Controls: Test/Retest Data for Systemic, Multi-Organ Analysis
The aim of the dataset was to develop a normative database for FDG-PET based on healthy controls, providing a reference for identifying voxel-level metabolic aberrations in cancer patients. This approach may uncover treatment-related changes beyond cancer diagnostics.
Read the related publication in Scientific Data.
Key Findings:
The dataset can be found in QUADRA_HC.
The MCWG Outreach Council invites you to submit announcements or information about papers, conferences, presentations or other events or news related to brain and molecular connectivity as well as any positions available or job opportunities that you wish to publicize and share with the community!
Please submit any material for consideration by the final day of each month using this form – thank you!

The MCWG is made up of four international and multidisciplinary councils dedicated to promoting molecular connectivity research via dissemination of methods, results, collaboration, and resource sharing (e.g. datasets, tools) within the scientific community. We encourage the neuroscientific community to take an integrative perspective in study of the brain connectome, where various methods including MRI-based techniques, electrophysiological tools, and molecular imaging advance our understanding of the brain. Please find fundamental questions outlined here: “Brain connectomics: time for a molecular imaging perspective?”
Our website can be found here. We also invite you to join the MCWG!
[MCWG] Molecular Connectivity Newsletter: November 2025

Thank you to everyone who joined us for October’s MCOS with Silvia Caminiti, PhD! Missed it? The recording is available here.
More details and key announcements below!
We hope you enjoy it!
Date: November 21st, 2025
Time: 15:00 CET, 09:00 EST (expected duration: 1.5h)
Registration: Please register here.
Title: Validating brain connectivity measures: integrating biological, statistical, and clinical evidence
Speaker: Alan Jasanoff, Department of Biological Engineering, MIT, USA
Andrea Hildebrandt, Department of Psychology, University of Oldenburg, Germany
Zhen-Qi Liu, Montreal Neurological Institute, McGill University, Canada
Matej Perovnik, Department of Neurology, University of Ljubljana University, Slovenia




Abstract: In this symposium, we will show how validation frameworks across biological, statistical and clinical dimensions help ground our interpretation of connectivity measures, establishing more reliable tools for both basic neuroscience and clinical applications. After a short introduction on the concept of validation by the organizers, Alan Jasanoff will tackle biological validation, showing how a novel genetically encoded activity reporter in animal models can inform the biological basis of fMRI brain connectivity. Second, Andrea Hildebrandt will show how a systematic characterization of the brain connectivity multiverse can provide valuable insights into assessment of statistical robustness across analytical choices as a prerequisite for enhancing replicability. Third, Zhen-Qi Liu will provide an overview of the properties of MR, MEG and PET brain connectivity measures, highlighting which metrics possess desirable statistical and biological properties. Last, Matej Perovnik will show a real-life example of systematic clinical validation based on PET brain networks..
The MCOS promotes rigor in research and resource sharing. We aim to hold MCOS every third Friday of the month, subject to change due to speaker availability. Please stay tuned for MCOS updates and reminders on social media! Thank you!
Each month, we will feature a member of the MCWG and have a brief Q&A!
This month please enjoy our highlight of Prof. Dr. Adriana Tavares, member of the MCWG Steering Committee.

Professor Adriana Tavares is a leading expert in translational molecular imaging at the University of Edinburgh, where she holds a Personal Chair in the Deanery of Clinical Sciences. She also serves as the Head of the Preclinical PET Facility within the Centre for Cardiovascular Science and is affiliated with Edinburgh Imaging. Her research focuses on the development of novel PET radiotracers and advanced image analysis techniques to enhance disease diagnosis, prognosis, and treatment monitoring. Her team is particularly engaged in creating molecular imaging biomarkers and kinetic modeling tools for whole-body PET studies. At the University of Edinburgh, she established the “PET is Wonderful” initiative, aimed at fostering collaboration and innovation in PET imaging research. She is an active member of the European Society of Molecular Imaging (ESMI) and co-chairs its STANDARD group. Additionally, she contributes to the Scottish Imaging Network: A Platform for Scientific Excellence (SINAPSE) and co-leads the UK PET Network’s preclinical subgroup. Professor Tavares has secured significant research funding, including a £1.93 million MRC award for developing the TSPO PET radiotracer [18F]LW223, aimed at detecting regional tissue inflammation. Her extensive publication record and leadership in the field underscore her commitment to advancing molecular imaging technologies for improved healthcare outcomes.
Prof Dr Adriana Tavares has graciously responded to our feature questionnaire:
What sparked your interest in molecular imaging or led you to focus on research in molecular imaging?
My first contact with Nuclear Medicine was when I started my BSc on that topic. I started my degree knowing I wanted to continue studying science, but I wasn’t quite sure how Nuclear Medicine worked and what could be done with radiotracers. As soon as my BSc classes started, I was completely hooked! I loved the idea of being able to look and quantify what is going on inside our cells in our bodies. This was my “Eureka” moment and, from that point onwards, I was sure that was the right career for me.
What is your role in the Molecular Connectivity Working Group, and what have you been contributing to or working on within the group?
I’m currently a member of the Molecular Connectivity Working Group and have been contributing with presentations on preclinical network analysis of PET datasets.
In what ways do you imagine molecular connectivity will advance our understanding of brain function?
I think it has already advanced our understanding of brain function over the years and, as new methodology comes to fore, I expect the impact to increase. For example, I envision a scenario where brain-body connectivity could inform how the multiple body systems are controlled by the brain and vice-versa.
What do you think are the most important challenges in current brain connectivity research, or which unsolved/underappreciated issues should the community address?
I think an important unaddressed challenge is how the brain connectivity tools could migrate from research into clinical practice. This has not really happened for most (if not all) of the tools and I think it would be worthwhile investigating and addressing the reasons underlying this observation, so the community can move forward from seeing the connectivity tools almost exclusively as research tools to more integrated clinical techniques.
What is your favorite mentoring memory—either a story about a mentor’s impact on you or your impact on a mentee?
One of my mentors once said that developing and communicating innovative tools can only be appreciated by peers and the wider community if well contextualised in terms of background (what exists now) and expected impact (what will change after implementing the new tool), otherwise the message will be lost in translation. At the time, they gave me an unexpected and hard to forget analogy: good science without context is like feeding pearls to pigs.
What scientist or scientific achievement do you most admire?
As a major radionuclide “fan” and a female scientist, I have great admiration for all scientific achievements by Maria Salomea Skłodowska-Curie.
📝 Multimodal metabolic and functional network signatures for diagnosis of parkinson’s disease: a PET/MR study
In this study, integrated PET/MR and Jensen-Shannon similarity estimation (JSSE) was employed to investigate metabolic/functional network dynamics in Parkinson’s disease (PD).
Read the full study in Scientific Reports.
Key Findings:
📝 Brain networks involved in cancer treatment response: insights from 18F-FDG PET scans
An exploratory retrospective study of two independent cohorts evaluated metabolic brain network scores from pre-treatment 18F-FDG PET scans and their ability to stratify good versus poor responders using ROC analysis (AUC). Longitudinal changes in network scores were assessed across follow-up, and progression-free survival (PFS) and overall survival (OS) analyses were performed in the melanoma cohort.
Read the full study preprint in Phys Med Bio.
Key Findings:
📝 Test-retest reproducibility of structural and proxy estimates of brain connectivity at rest
This study compared test-retest reproducibility of group-level structural connectivity, functional connectivity, intersubject covariance of regional gray matter volume (GMVcov), and intersubject covariance of regional [18F]Fluorodeoxyglucose uptake (FDGcov) in the same 55 healthy subjects at rest using a simultaneous PET/MRI acquisition protocol.
Read the full study preprint in NeuroImage.
Key Findings:
The MCWG Outreach Council invites you to submit announcements or information about papers, conferences, presentations or other events or news related to brain and molecular connectivity as well as any positions available or job opportunities that you wish to publicize and share with the community!
Please submit any material for consideration by the final day of each month using this form – thank you!

The MCWG is made up of four international and multidisciplinary councils dedicated to promoting molecular connectivity research via dissemination of methods, results, collaboration, and resource sharing (e.g. datasets, tools) within the scientific community. We encourage the neuroscientific community to take an integrative perspective in study of the brain connectome, where various methods including MRI-based techniques, electrophysiological tools, and molecular imaging advance our understanding of the brain. Please find fundamental questions outlined here: “Brain connectomics: time for a molecular imaging perspective?”
Our website can be found here. We also invite you to join the MCWG!
[MCWG] Molecular Connectivity Newsletter: October 2025

Thank you to everyone who joined us for September’s MCOS with Jingyuan Chen PhD! Missed it? The recording is available here.
More details and key announcements below!
We hope you enjoy it!
Date: October 17th, 2025
Time: 15:00 CEST, 09:00 EDT
Registration: Please register here.
Title: Metabolic connectivity features in Alzheimer’s disease
Speaker: Silvia Paola Caminiti
Abstract: Emerging evidence highlights that connectivity alterations in Alzheimer’s disease (AD) include not only hypo-connectivity, but also hyper-connectivity, particularly in limbic and cerebellar regions during the prodromal and early dementia stages. In a recent longitudinal multimodal study, it was demonstrated that hyper-connectivity is detectable with FDG-PET metabolic connectivity and is not merely compensatory. These findings converge with other recent reports showing that medial temporal and anterior-temporal hyper-connectivity mediates amyloid-related tau accumulation and accelerates neurodegeneration. Altogether, converging data suggest that hyper-connectivity, far from being a benign or compensatory feature, may actively drive AD progression by increasing metabolic burden and synaptic inefficiency. Understanding this dual landscape of hypo- and hyper-connectivity offers new insights into AD pathophysiology and may open perspectives for network-based biomarkers of disease progression.

Dr. Caminiti is an Assistant Professor at the Department of Brain and Behavioral Sciences at the University of Pavia in Italy. With a background in neuroimaging and neuroscience, she is dedicated to advancing our understanding of the intricacies of the brain and its functioning. Her primary research focus revolves around aging and neurodegenerative diseases. The goal of her research program is to examine selective neuronal vulnerability in Alzheimer’s and Parkinson’s disease using multimodal neuroimaging including PET and MRI.
The MCOS promotes rigor in research and resource sharing. We aim to hold MCOS every third Friday of the month, subject to change due to speaker availability. Please stay tuned for MCOS updates and reminders on social media! Thank you!
📝 Building multivariate molecular imaging brain atlases using the NeuroMark PET independent component analysis framework
Eierud et al. introduce the NeuroMark PET approach, utilizing spatially constrained independent component analysis (ICA) to define overlapping regions that may reflect the brain’s molecular architecture.
Read the full study in Aperture Neuro.
Key Findings:
📝 Glucose Metabolism echoes Long-Range Temporal Correlations in the Human Brain
In this study, Facca and colleagues explored the implications for individual-level metabolic regulation by integrating resting-state functional Magnetic Resonance Imaging (fMRI) and dynamic [18F]FDG Positron Emission Tomography (PET) data acquired from the same cohort of participants.
Read the full study preprint in bioRxiv.
Key Findings:
The MCWG Outreach Council invites you to submit announcements or information about papers, conferences, presentations or other events or news related to brain and molecular connectivity as well as any positions available or job opportunities that you wish to publicize and share with the community!
Please submit any material for consideration by the final day of each month using this form – thank you!

The MCWG is made up of four international and multidisciplinary councils dedicated to promoting molecular connectivity research via dissemination of methods, results, collaboration, and resource sharing (e.g. datasets, tools) within the scientific community. We encourage the neuroscientific community to take an integrative perspective in study of the brain connectome, where various methods including MRI-based techniques, electrophysiological tools, and molecular imaging advance our understanding of the brain. Please find fundamental questions outlined here: “Brain connectomics: time for a molecular imaging perspective?”
Our website can be found here. We also invite you to join the MCWG!
[MCWG] Molecular Connectivity Newsletter: September 2025

More details and key announcements below!
We hope you enjoy it!
Date: September 19th, 2025
Time: 15:00 CEST, 09:00 EDT
Registration: Please register here.
Title: Simultaneous EEG-PET-MRI identifies temporally coupled, spatially structured hemodynamic and metabolic dynamics across wakefulness and NREM sleep.
Speaker: Jingyuan Chen
Abstract: Sleep induces profound changes in cerebral hemodynamics and metabolism, yet how these processes unfold across wakefulness and sleep, as well as their spatiotemporal dependence remains largely unknown. In this talk, Dr. Chen will present her recent work integrating functional PET (fPET)-FDG with simultaneous EEG-fMRI to track physiological dynamics across the sleep–wake cycle. Findings show that global hemodynamic and metabolic signals are tightly coupled during the descent into NREM sleep, both closely tracking EEG arousal dynamics. There were identified two distinct network patterns in NREM sleep. These findings shed light on how sleep can diminish awareness while preserving sensory responsiveness, revealing a complex, alternating balance of hemodynamic and metabolic processes in the sleeping brain. This work also highlights the potential of EEG–PET–MRI for probing the neuro-hemo-metabolic underpinnings of cognition and arousal in humans.

Dr. Chen is an Assistant Professor at the Martinos Center for Biomedical Imaging at Massachusetts General Hospital and Harvard Medical School. She received her Ph.D. from Stanford University, with a major in Electrical Engineering and a minor in Statistics. Following graduation, she pursued further training in neuroimaging as a postdoctoral fellow at the Martinos Center. Her lab advances multimodal dynamic functional imaging techniques and computational approaches to investigate the biophysical and molecular mechanisms underlying intrinsic brain activity, as well as the neurobiological consequences of cognition, arousal, and disease.
Date: October 17th, 2025
Time: 15:00 CEST, 09:00 EDT
Registration: Please register here.
Title: Metabolic connectivity features in Alzheimer’s disease.
Speaker: Silvia Paola Caminiti
The MCOS promotes rigor in research and resource sharing. We aim to hold MCOS every third Friday of the month, subject to change due to speaker availability. Please stay tuned for MCOS updates and reminders on social media! Thank you!
📝 On the analysis of functional PET (fPET)-FDG: baseline mischaracterization can introduce artifactual metabolic (de)activations
Coursey et al. investigated how inaccuracies in modeling baseline FDG uptake can introduce artifactual patterns to detrended time-activity curve (TAC) residuals, potentially introducing spurious (de)activations to general linear model (GLM) analyses.
Read the full study in Imaging Neuroscience.
Key Findings:
📝 Stable brain PET metabolic networks using a multiple sampling scheme
In this study, Shu and colleagues used brain [18F]fluoro-2-deoxyglucose PET data from 1,227 individuals in the Alzheimer’s disease (AD) continuum from the Alzheimer’s Disease Neuroimaging Initiative cohort to develop a novel method for constructing stable metabolic brain networks that are resilient to spurious data points.
Read the full study preprint in Network Neuroscience.
Key Findings:
📝 Comparative evaluation of graph construction methods for individual brain metabolic network from FDG-PET images: an ADNI study in healthy subjects
Graph theory is emergently applied to model brain networks at individual level; however, the choice of graph construction method can significantly impact analytical outcomes. In this study, methods for building individual graphs from FDG-PET images in healthy control subjects were systematically evaluated and compared by Tuan et al.
Read the full study preprint in Eur J Nucl Med Mol Imaging.
Key Findings:
The MCWG Outreach Council invites you to submit announcements or information about papers, conferences, presentations or other events or news related to brain and molecular connectivity as well as any positions available or job opportunities that you wish to publicize and share with the community!
Please submit any material for consideration by the final day of each month using this form – thank you!

The MCWG is made up of four international and multidisciplinary councils dedicated to promoting molecular connectivity research via dissemination of methods, results, collaboration, and resource sharing (e.g. datasets, tools) within the scientific community. We encourage the neuroscientific community to take an integrative perspective in study of the brain connectome, where various methods including MRI-based techniques, electrophysiological tools, and molecular imaging advance our understanding of the brain. Please find fundamental questions outlined here: “Brain connectomics: time for a molecular imaging perspective?”
Our website can be found here. We also invite you to join the MCWG!
[MCWG] Molecular Connectivity Newsletter: August 2025

Greetings from the MCWG! Wishing you a bright and restful summer!
MCWG members were present at the Brain and Brain PET 2025 conference in Seoul and at the Annual Meeting of the Organization for Human Brain Mapping (OHBM) 2025 conference in Brisbane.
We’ve put together some highlights and presentations showcasing recent advances in molecular connectivity.
We hope you enjoy it!

🧠 Symposium: Current advances in molecular connectivity
Tommaso Volpi, PhD


Sharna Jamadar, PhD


Kristina Herfert, PhD


📝 Featured Abstracts
Metabolic Connectivity From Dynamic [18F]FDG PET Reveals Task-Evoked Brain Network Architecture
Authors: Giulia Vallini, Murray Bruce Reed, Sebastian Klug, Godber Mathis Godbersen, Alessandra Bertoldo, Rupert Lanzenberger, and Andreas Hahn
Exercise Partially Restores Altered Metabolic Connectivity in Parkinson’s Disease
Authors: Connor W. J. Bevington, Erik Reimers, Jess Mckenzie, Sahib Dhaliwal, Teresa Liu-ambrose, A. Jon Stoessl, and Vesna Sossi
Desynchronization Within the Striatal Network in Schizophrenia: Preliminary Results of an FDOPA-PET Study
Authors: Yifan Mayr, Julia Schulz, Melissa Thalhammer, Mattia Veronese, Christian Sorg, Felix Brandl, and Igor Yakushev
🧠 Keynote Presentation
Sharna Jamadar, PhD
Glucodynamics and the metabolic network of the human brain
🧠 Symposium: Relationship between Brain Energy Consumption and Brain Organizations
Organizer: Xin Di


Featured Talks:

🧠 Symposium: Validating brain connectivity measures: integrating biological, statistical, and clinical evidence
Organizers: Chris Habeck, Arianna Sala, Daniel Talmasov, Vesna Sossi
Moderators: James Pang, Felix Hoffstaedter
Featured Talks:


🧠 Oral Presentation
Mengyuan Liu
Unveiling the biological substrates of glucose metabolic covariance in the human brain
📝 Featured Abstracts
Single-subject network analysis of FDOPA PET in Parkinson’s disease and Schizophrenia
Authors: Mario Severino, Julia Schubert, Giovanna Nordio, Alessio Giacomel, Rubaida Easmin, Nick P. Lao-Kaim, Pierluigi Selvaggi, Zhilei Xu, Joana B. Pereira, Sameer Jahuar, Paola Piccini, Oliver Howes, Federico Turkheimer, Mattia Veronese
Brain functional-metabolic relationships:rs-fMRI/dynamic [18F]FDG-PET multivariate integration
Authors: Claudia Tarricone, Giulia Vallini, Giorgia Baron, Erica Silvestri, Tommaso Volpi, Andrei G. Vlassenko, Manu S. Goyal, Alessandra Bertoldo
📝 Comparing intra- and inter-individual correlational brain connectivity from functional and structural neuroimaging data
This study examined intra-individual correlations of structural and functional brain measures across an extended time span. The study focused on intra-individual correlations, with the aim to minimize individual differences and investigate how aging and state-like effects contribute to brain connectivity patterns. Intra-individual correlations with inter-individual correlations were compared. Read the full study in Brain Struct Funct.
Key Findings:
📝 Brain glucodynamic variability is an essential feature of the metabolism-cognition relationship
In this study the temporal variability in glucodynamics, or time-varying glucose use, related to cognition was evaluated in 35 younger and 43 older adults. Read the full study preprint in bioRxiv.
Key Findings:
📝 Unveiling functional-metabolic synergy in the healthy brain: multivariate integration of dynamic [18F]FDG-PET and resting-state fMRI
This study employed a multivariate Partial Least Squares Correlation (PLSC) to investigate the functional-metabolic relationship at both nodal and network level with dynamic [18F]FDG-PET data, generated within-individual metabolic connectivity (MC) networks, and functional connectivity (FC) derived from fMRI data. Read the full study in bioRxiv.
Key Findings:
The MCOS promotes rigor in research and resource sharing. We aim to hold MCOS every third Friday of the month, subject to change due to speaker availability.
Please stay tuned for MCOS updates and reminders on social media!
Thank you!
The MCWG Outreach Council invites you to submit announcements or information about papers, conferences, presentations or other events or news related to brain and molecular connectivity as well as any positions available or job opportunities that you wish to publicize and share with the community!
Please submit any material for consideration by the final day of each month using this form – thank you!

The MCWG is made up of four international and multidisciplinary councils dedicated to promoting molecular connectivity research via dissemination of methods, results, collaboration, and resource sharing (e.g. datasets, tools) within the scientific community. We encourage the neuroscientific community to take an integrative perspective in study of the brain connectome, where various methods including MRI-based techniques, electrophysiological tools, and molecular imaging advance our understanding of the brain. Please find fundamental questions outlined here: “Brain connectomics: time for a molecular imaging perspective?”
Our website can be found here. We also invite you to join the MCWG!
[MCWG] Molecular Connectivity Newsletter: June 2025

Greetings from the MCWG!
Thank you to everyone who joined us for May’s MCOS with Simon Eickhoff, PhD! Missed it? The recording is available here.
June 24-28, 2025
Heading to the Annual Meeting of the Organization for Human Brain Mapping (OHBM) 2025 in Brisbane?
We’ve put together a list of events and presentations showcasing recent advances in molecular connectivity.
We hope to see you there!
🧠 Keynote Presentation
Sharna Jamadar, PhD
Glucodynamics and the metabolic network of the human brain
🗓 June 28 | ⏰ 10:30–11:15
🧠 Symposium: Relationship between Brain Energy Consumption and Brain Organizations
Organizer: Xin Di
🗓 June 27 | ⏰ 9:00–10:15
Featured Talks:
🧠 Symposium: Validating brain connectivity measures: integrating biological, statistical, and clinical evidence
Organizers: Chris Habeck, Arianna Sala, Daniel Talmasov, Vesna Sossi
Moderators: James Pang, Felix Hoffstaedter
🗓 June 27 | ⏰ 15:45–17:00
Featured Talks:
🧠 Oral Presentation
Mengyuan Liu
Unveiling the biological substrates of glucose metabolic covariance in the human brain
🗓 June 28 | ⏰ 11:42–11:54
📝 Featured Abstract
Power and Discordance of Exercise Intervention on Parkinson’s Disease fMRI Functional Connectivity
Authors: Erik Reimers, Connor W. J. Bevington, Jess McKenzie, Sahib Dhaliwal, A. Jon Stoessl, Vesna Sossi
2nd – 4th July 2025 • Edinburgh, UK
Featuring two days Education Workshop and one day Scientific meeting on PET. Highlights include an educational session on Network analysis and radiomics in total-body PET imaging by Dr. Thomas Beyer. More information about the event click here.
📝 Glucose metabolism in hyper-connected regions predicts neurodegeneration and speed of conversion in Alzheimer’s disease
Longitudinal analysis of brain metabolic connectivity on neurodegeneration in Alzheimer’s disease. Read the full study in Eur J Nucl Med Mol Imaging.
Key Findings:
📝 Innate [18F]Fluorodeoxyglucose PET bone networks of lung cancer patients predict survival
Newly proposed network analysis performs better than conventional SUV PET analysis, and should be considered as a prognostication tool in the management of NSCLC patients. Read the full study in Eur J Nucl Med Mol Imaging.
Key Findings:
📝 Mapping serotonergic dynamics using drug-modulated molecular connectivity in rats
Brain molecular connectivity was measured using combined serotonin PET and resting-state fMRI providing insights into the effects of ecstasy administration. Read the full study in Neuroscience eLife.
Key Findings:
📝 Multiplex connectomics reveal altered networks in frontotemporal dementia: A multisite study
A recent multisite study employed an innovative multiplex connectomics framework integrating cortical thickness and FDG-PET data to uncover how brain networks are disrupted in frontotemporal dementia (FTD). An employed dual-layer approach provides a new lens to understand neurodegenerative disease mechanisms. Read the full study in Network Neuroscience.
Key Findings:
📝 Single-Subject Network Analysis of FDOPA PET in Parkinson’s Disease and Psychosis Spectrum
In this study, a statistical framework was proposed combining molecular imaging data with perturbation covariance analysis to construct single-subject networks and investigate individual patterns of molecular alterations. Mapping molecular imaging networks presents a new and powerful method for characterizing individualized disease trajectories as well as for evaluating treatment effectiveness. Read the full study in Hum Brain Mapp.
Key Findings:
📝 Fully automated estimation of fMRI-guided SPECT brain networks and their functional network connectivity in schizophrenia patients versus controls: A NeuroMark ICA approach
In this study, a fully automated, spatially constrained ICA (sc-ICA) approach to evaluate functional network connectivity profiles in SPECT data using the NeuroMark pipeline was presented. The expression of brain networks along with the whole brain SPECT connectome to evaluate neuroimaging links to schizophrenia were evaluated. Read the full study in Aperture Neuro.
Key Findings:
The MCOS promotes rigor in research and resource sharing. We aim to hold MCOS every third Friday of the month, subject to change due to speaker availability.
Please stay tuned for MCOS updates and reminders on social media!
Thank you!
The MCWG Outreach Council invites you to submit announcements or information about papers, conferences, presentations or other events or news related to brain and molecular connectivity as well as any positions available or job opportunities that you wish to publicize and share with the community!
Please submit any material for consideration by the final day of each month using this form – thank you!

The MCWG is made up of four international and multidisciplinary councils dedicated to promoting molecular connectivity research via dissemination of methods, results, collaboration, and resource sharing (e.g. datasets, tools) within the scientific community. We encourage the neuroscientific community to take an integrative perspective in study of the brain connectome, where various methods including MRI-based techniques, electrophysiological tools, and molecular imaging advance our understanding of the brain. Please find fundamental questions outlined here: “Brain connectomics: time for a molecular imaging perspective?”
Our website can be found here. We also invite you to join the MCWG!
[MCWG] Molecular Connectivity Newsletter: May 2025

Greetings from the MCWG!
Thank you to everyone who joined us for April’s MCOS with Mattia Veronese PhD & Mario Severino! Missed it? The recording is available here.
Please join us for the May MCOS featuring a talk by Simon Eickhoff PhD, Heinrich-Heine University. More details and key announcements below!
Date: May 23rd, 2025
Time: 15:00 CET, 09:00 EDT
Registration: Please register here.
Title: Great expectations but a bumpy road: Caveats in neuroimaging analyses and modelling.
Speaker: Simon Eickhoff, PhD
Abstract: The increasing availability of large cohorts and tools for multivariate statistical learning, allowing the prediction of individual cognitive or clinical phenotypes in new subjects, started a revolution towards neuroimaging applications in precision medicine. While the field has enjoyed a lot of enthusiasm recently, the road towards translation and re-life applications may be considerably more challenging than often acknowledged. Following a short overview on the motivation and perspectives, the major part of my talk will focus on several critical yet often underappreciated challenges for such endeavors. These include on the one hand technical and biological aspects that may undermine the validity of prediction results, due to the inherent low-dimensional structure of biological variability. On the other hand, ethical, legal and societal aspects will ultimately shape practical adaptation but need stronger consideration in the development of new pipelines if these are to move beyond proof-of-concept work.

Simon Eickhoff is a full professor and chair of the Institute for Systems Neuroscience at the Heinrich-Heine University in Düsseldorf and the director of the Institute of Neuroscience and Medicine (INM-7, Brain and Behavior) at the Forschungszentrum Jülich. He is furthermore a visiting professor at the Chinese Academy of Science Institute of Automation. Working at the interface between neuroanatomy, data-science and brain medicine, he aims to obtain a more detailed characterization of the organization of the human brain and its inter-individual variability in order to better understand its changes in advanced age as well as neurological and psychiatric disorders. This goal is pursued by the development and application of novel analysis tools and approaches for large-scale, multi-modal analysis of brain structure, function and connectivity as well as by machine-learning for single subject prediction of cognitive and socio-affective traits and ultimately precision medicine.
The MCOS promotes rigor in research and resource sharing. We aim to hold MCOS every third Friday of the month, subject to change due to speaker availability. Please stay tuned for MCOS updates and reminders on social media! Thank you!
Heading to Brain & Brain PET 2025 in Seoul (June 1–4)?
We’ve compiled a list of events and presentations at the meeting that highlight recent developments in molecular connectivity. We hope to see you there!
🗓 June 1 | 14:00–17:30
Moderator: Mark Lubberink
Speakers: Igor Yakushev, Giulia Vallini, Sharna Jamadar, Murray Reed
🗓 June 4 | 11:00–12:30
Organizer: Sharna Jamadar
Featured Talks:
Exercise Effects on Abnormal Discordance Between Glucose Metabolism and Cerebral Blood Flow in Parkinson’s Disease
Connor W. J. Bevington, Jess McKenzie, Sahib Dhaliwal, Teresa Liu-Ambrose, A. Jon Stoessl, Vesna Sossi
Exercise Partially Restores Altered Metabolic Connectivity in Parkinson’s Disease
Connor W. J. Bevington, Erik Reimers, Jess McKenzie, Sahib Dhaliwal, Teresa Liu-Ambrose, A. Jon Stoessl, Vesna Sossi
Metabolic Connectivity Alterations in Post-COVID Condition
Patients with post-COVID condition (PCC) exhibit lasting reorganization of brain metabolic connectivity, according to our recent study using ¹⁸F-FDG PET. These findings shed new light on the long-term effects of COVID-19 on brain function.
Read the full study in EJNMMI Research
The MCWG Outreach Council invites you to submit announcements or information about papers, conferences, presentations or other events or news related to brain and molecular connectivity as well as any positions available or job opportunities that you wish to publicize and share with the community!
Please submit any material for consideration by the final day of each month using this form – thank you!

The MCWG is made up of four international and multidisciplinary councils dedicated to promoting molecular connectivity research via dissemination of methods, results, collaboration, and resource sharing (e.g. datasets, tools) within the scientific community. We encourage the neuroscientific community to take an integrative perspective in study of the brain connectome, where various methods including MRI-based techniques, electrophysiological tools, and molecular imaging advance our understanding of the brain. Please find fundamental questions outlined here: “Brain connectomics: time for a molecular imaging perspective?”
Our website can be found here. We also invite you to join the MCWG!
[MCWG] Molecular Connectivity Newsletter: April 2025

Greetings to all!
We wish to thank all who attended MCOS in March with Xin Di, PhD! The recording from his talk can be found here. Please join us for the April MCOS featuring a talk by Mattia Veronese, PhD & Mario Severino, University of Padua, please find more information and additional announcements below!
We are pleased to announce the next MCOS talk featuring Mattia Veronese, PhD & Mario Severino!
Date: April 11th, 2025
Time: 15:00 CET, 09:00 EDT
Title: Molecular Connectivity in Neurotransmission.
Please join us for this 30 minute presentation to be followed by discussion (~25 minutes).
Please register here.
Abstract: PET and SPECT are fundamental tools for studying neurotransmission in living systems. Traditionally, these imaging modalities are analysed by focusing on individual regions or voxels independently. However, recent interest has shifted toward molecular connectivity approaches, which offer a fresh perspective on the brain’s complex organization by examining interactions between brain regions. This talk will provide a comprehensive review of molecular connectivity studies in neurotransmission, highlighting their expanding applications, advantages over traditional methods, and contributions to advancing neuroscience.

Dr. Mattia Veronese is Associate Professor in Biomedical Engineering at the Department of Information Engineering, University of Padua and Honorary Senior Lecturer in Neuroimaging at King’s College London. He is a biomedical engineer by training and holds a PhD in PET kinetic modelling. His main research interest is related to the development and validation of molecular neuroimaging biomarkers and to their use for drug development and precision medicine.
Click here for more information about him.

Mario Severino is a PhD student in Bioengineering at the Department of Information Engineering, University of Padua. He holds a Master’s degree in Bioengineering. His primary research focus is on applying PET molecular connectivity and network science approaches, aiming to establish these methods as precision medicine biomarkers.
The MCOS promotes rigor in research and resource sharing. We aim to hold MCOS every third Friday of the month, subject to change due to speaker availability.
Upcoming MCOS Speakers
The scoping review from the MCWG is finally out in Imaging Neuroscience!
Molecular connectivity studies in neurotransmission: a scoping review
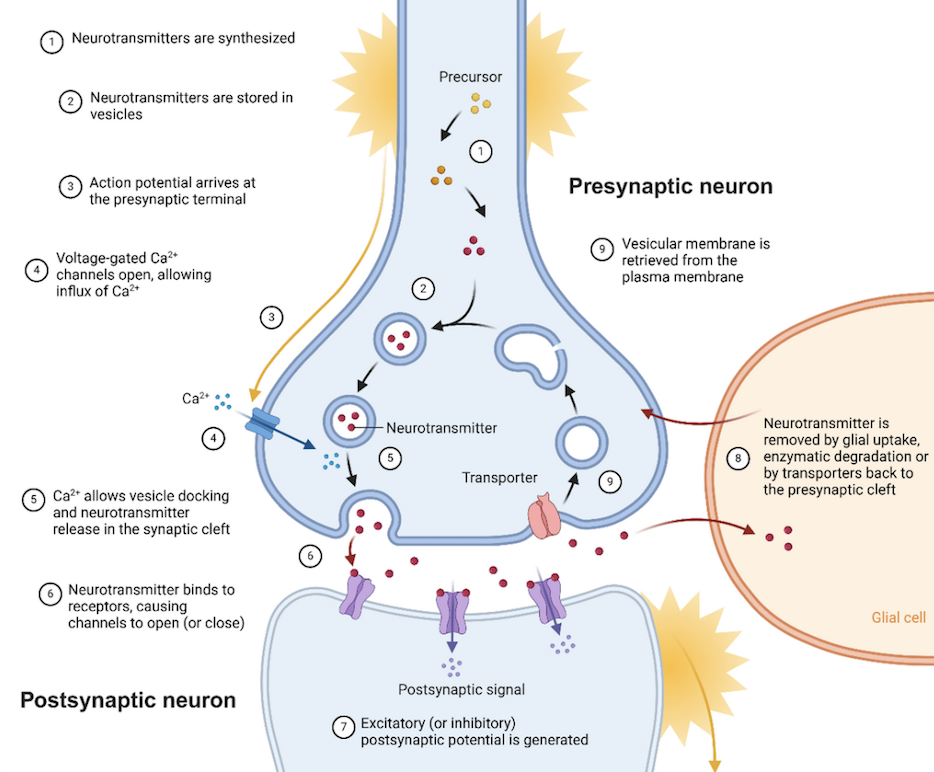
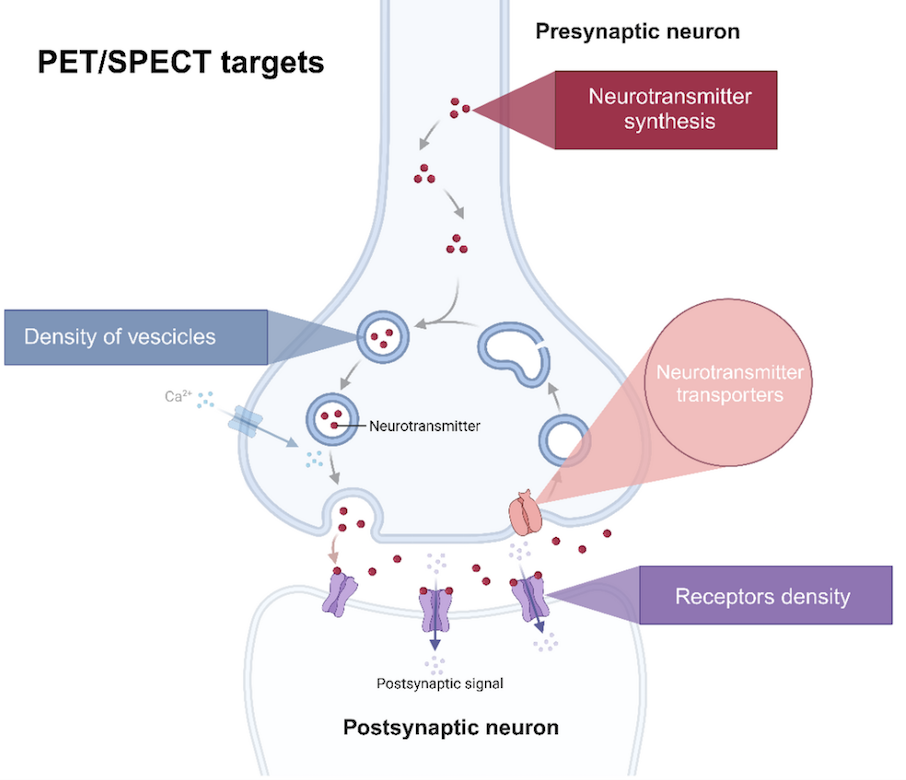
Abstract: Positron emission tomography (PET) and single photon emission computed tomography (SPECT) are essential molecular imaging tools for the in vivo investigation of neurotransmission. Traditionally, PET and SPECT images are analysed in a univariate manner, testing for changes in radiotracer binding in regions or voxels of interest independently of each other. Over the past decade, there has been an increasing interest in the so-called molecular connectivity approach that captures relationships of molecular imaging measures in different brain regions. Targeting these inter-regional interactions within a neuroreceptor system may allow to better understand complex brain functions. In this article, we provide a comprehensive review of molecular connectivity studies in the field of neurotransmission. We examine the expanding use of molecular connectivity approaches, highlighting their applications, advantages over traditional methods, and contributions to advancing neuroscientific knowledge. A systematic search in three bibliographic databases MEDLINE, EMBASE and Scopus on July 14, 2023, was conducted. A second search was rerun on April 4, 2024. Molecular imaging studies examining functional interactions across brain regions were included based on predefined inclusion and exclusion criteria. Thirty-nine studies were included in the scoping review. Studies were categorised based on the primary neurotransmitter system being targeted: dopamine, serotonin, opioid, muscarinic, glutamate and synaptic density. The most investigated system was the dopaminergic and the most investigated disease was Parkinson’s disease (PD). This review highlighted the diverse applications and methodologies in molecular connectivity research, particularly for neurodegenerative diseases and psychiatric disorders. Molecular connectivity research offers significant advantages over traditional methods, providing deeper insights into brain function and disease mechanisms. As the field continues to evolve, embracing these advanced methodologies will be essential to understand the complexities of the human brain and improve the robustness and applicability of research findings in clinical settings.
Please stay tuned for MCOS updates and reminders on social media! Thank you!
The MCWG Outreach Council invites you to submit announcements or information about papers, conferences, presentations or other events or news related to brain and molecular connectivity as well as any positions available or job opportunities that you wish to publicize and share with the community!
Please submit any material for consideration by the final day of each month using this form – thank you!

The MCWG is made up of four international and multidisciplinary councils dedicated to promoting molecular connectivity research via dissemination of methods, results, collaboration, and resource sharing (e.g. datasets, tools) within the scientific community. We encourage the neuroscientific community to take an integrative perspective in study of the brain connectome, where various methods including MRI-based techniques, electrophysiological tools, and molecular imaging advance our understanding of the brain. Please find fundamental questions outlined here: “Brain connectomics: time for a molecular imaging perspective?”
Our website can be found here. We also invite you to join the MCWG!
[MCWG] Molecular Connectivity Newsletter: March 2025

Greetings to all!
We wish to thank all who attended MCOS in Feburary with Kristina Herfert, PhD! The recording from her talk can be found here. Please join us for the March MCOS featuring a talk by Xin Di, PhD, New Jersey Institute of Technology, please find more information and additional announcements below!
We are pleased to announce the next MCOS talk featuring Xin Di, PhD, New Jersey Institute of Technology!
Date: March 21st, 2025
Time: 15:00 CET, 10:00 EDT
Title: Comparing Intra- and Inter-individual Correlational Brain Connectivity from Functional and Structural Neuroimaging Data.
Please join us for this 30 minute presentation to be followed by discussion (~25 minutes).
Please register here.
Abstract: Many neuroimaging modalities, such as PET and structural MRI, typically assess connectivity using inter-individual correlations, although these measures may be affected by several factors. To clarify these influences, we compared intra- and inter-individual correlations using identical data types. Our analysis of two distinct datasets revealed that long-term age effects contribute to correlations in brain structure, though they bear limited resemblance to functional connectivity. In contrast, both intra- and inter-individual correlations in functional measures (PET and ReHo) strongly reflected functional connectivity, underscoring the importance of state-like brain activity. These findings highlight the need to account for such influences when interpreting inter-individual connectivity measures.

Dr. Xin Di is a Research Assistant Professor in the Department of Biomedical Engineering at the New Jersey Institute of Technology. With a PhD in psychology, his research focuses on modeling brain connectivity using multi-modal neuroimaging data, including fMRI, PET, structural MRI, and fNIRS. He also investigates brain alterations associated with autism spectrum disorder.
Please find further information about Dr. Di here!
The MCOS promotes rigor in research and resource sharing. We aim to hold MCOS every third Friday of the month, subject to change due to speaker availability.
Upcoming MCOS Speakers
Announcements related to OHBM 2025 in June
In very exciting news, members of the MCWG will hold two separate symposia during the upcoming OHBM conference in June! Congratulations to all organizers and presenters! We wish to congratulate MCWG member, Sharna Jamadar, PhD who has been invited to give a keynote lecture at OHBM 2025. Dr. Jamadar will discuss work in her lab related to molecular connectivity, so please stay tuned for more information as OHBM approaches!
Christian Habeck, PhD Columbia University, Organizer, Arianna Sala, PhD University of Liège, Co-Organizer, Vesna Sossi, PhD, University of British Columbia, Co-Organizer, Daniel Talmasov, MD, Columbia University, Co-Organizer
Presentations
Xin Di, PhD, New Jersey Institute of Technology, Organizer
Presentations
A recent publication from the group led by MCWG member, Sharna Jamadar, PhD.
Deery et al., 2025, Metabolic connectivity has greater predictive utility for age and cognition than functional connectivity
“One point is that we feel that the results underscore the value of examining metabolic connectivity in addition to fMRI functional connectivity to understand the network dynamics of the brain. We did not set out to ‘prove’ that MC is somehow better than FC, rather to continue our story that MC provides important unique insight into the brain”. – Sharna Jamadar, PhD
Please stay tuned for MCOS updates and reminders on social media! Thank you!
We hope you will take a moment to read the MCWG authored preprint!
Molecular connectivity studies in neurotransmission: a scoping review


Purpose: Positron emission tomography (PET) and single photon emission computed tomography (SPECT) are essential molecular imaging tools for the in vivo investigation of neurotransmission. Traditionally, PET and SPECT images are analysed in a univariate manner, testing for changes in radiotracer binding in regions or voxels of interest independently of each other. Over the past decade, there has been an increasing interest in the so-called molecular connectivity approach that captures relationships of molecular imaging measures in different brain regions. Targeting these inter-regional interactions within a neuroreceptor system may allow to better understand complex brain functions. In this article, we provide a comprehensive review of molecular connectivity studies in the field of neurotransmission. We examine the expanding use of molecular connectivity approaches, highlighting their applications, advantages over traditional methods, and contributions to advancing neuroscientific knowledge.
The MCWG Outreach Council invites you to submit announcements or information about papers, conferences, presentations or other events or news related to brain and molecular connectivity as well as any positions available or job opportunities that you wish to publicize and share with the community!
Please submit any material for consideration by the final day of each month using this form – thank you!

The MCWG is made up of four international and multidisciplinary councils dedicated to promoting molecular connectivity research via dissemination of methods, results, collaboration, and resource sharing (e.g. datasets, tools) within the scientific community. We encourage the neuroscientific community to take an integrative perspective in study of the brain connectome, where various methods including MRI-based techniques, electrophysiological tools, and molecular imaging advance our understanding of the brain. Please find fundamental questions outlined here: “Brain connectomics: time for a molecular imaging perspective?”
Our website can be found here. We also invite you to join the MCWG!
[MCWG] Molecular Connectivity Newsletter: February 2025

Greetings to all!
We wish to thank all who attended our January MCOS with Mary C. Catanese, PhD! Please join us for the February MCOS with Kristina Herfert, PhD, please find more information in the announcement below! A new open postdoctoral position is also featured below!
We are pleased to announce the next MCOS talk featuring Kristina Herfert, PhD, University Hospital of Tuebingen!
Date: February 21st, 2025
Time: 15:00 CET, 9:00 EST
Title: Mapping brain function and connectivity in rodents using small animal PET/MRI.
Please join us for this 30 minute presentation to be followed by discussion (~25 minutes).
Please register here.
Abstract: Simultaneous small-animal PET/MRI integrates molecular and hemodynamic measures to offer deeper insights into brain function and connectivity. In our recent study, we combined functional PET (fPET) and fMRI to investigate the nigrostriatal pathway in rats under optogenetic stimulation, revealing pronounced inhibition in specific downstream regions rather than widespread activation. This indicates an unexpected inhibitory regulation within this key motor circuit. In another study, we compared molecular and functional connectivity measures following MDMA administration in rats. By pairing PET-based imaging of molecular targets with BOLD fMRI, we uncovered complementary insights into how neurotransmitter systems influence large-scale networks. These results underscore the power of fPET/fMRI to detect subtle changes in neuronal and molecular processes, highlighting the added value of multimodal imaging in shaping our understanding of brain function and informing both basic neuroscience and clinical research.

Prof. Dr. rer. nat. Kristina Herfert is Associate Professor for Functional and Metabolic Brain Imaging at the Werner Siemens Imaging Center, University of Tübingen, Germany. A trained neuroscientist, she specializes in in vivo PET quantification of dopaminergic and serotonergic receptors, PET tracer development for brain protein aggregates, and simultaneous small-animal PET/fMRI to study molecular and functional networks. After completing her doctorate under Prof. B.J. Pichler, she was awarded a Research Fellowship in Prof. Feng Zhang’s lab at the Broad Institute of MIT and Harvard. Professor Herfert’s research uniquely bridges gene editing approaches with neuroimaging to understand the brain’s molecular, cellular, and network-level functions.
Please find further information about Dr. Herfert here!
The MCOS promotes rigor in research and resource sharing. We aim to hold MCOS every third Friday of the month, subject to change due to speaker availability.
Upcoming MCOS Speakers (following a brief hiatus in March):
Please stay tuned for MCOS updates and reminders on social media!
Post-doctoral position: Dr. Joana Pereira at Karolinska Institute, Stockholm, Sweden.
The Pereira lab combines brain imaging with different machine learning algorithms, including deep learning, genetic as well as proteomics data to understand the pathological processes that occur in aging and neurodegenerative diseases.
This project will be carried out under Dr. Pereira’s supervision, and will involve analyzing cross-sectional and longitudinal imaging data, combining clinical measures and cerebrospinal fluid biomarkers, as well as writing manuscripts and presenting results in international conferences. The Postdoctoral fellow will have the opportunity to work in a friendly and dynamic environment, be involved in many projects, collaborate with group members on different studies, learn new skills, interact with researchers with different backgrounds and from other universities, analyze large datasets from well-characterized cohorts, and develop a wide network of international collaborations.
The ideal candidate should have the following qualities:
The Postdoctoral studies will be performed for 1 year, with possible extension for an additional year. There are also possibilities for a longer appointment after this 2-year period.
To apply please go to: https://ki.varbi.com/en/what:job/jobID:793528/ (Deadline: 20.Feb.2025)
Thank you!
We hope you will take a moment to read the MCWG authored preprint!
Molecular connectivity studies in neurotransmission: a scoping review


Purpose: Positron emission tomography (PET) and single photon emission computed tomography (SPECT) are essential molecular imaging tools for the in vivo investigation of neurotransmission. Traditionally, PET and SPECT images are analysed in a univariate manner, testing for changes in radiotracer binding in regions or voxels of interest independently of each other. Over the past decade, there has been an increasing interest in the so-called molecular connectivity approach that captures relationships of molecular imaging measures in different brain regions. Targeting these inter-regional interactions within a neuroreceptor system may allow to better understand complex brain functions. In this article, we provide a comprehensive review of molecular connectivity studies in the field of neurotransmission. We examine the expanding use of molecular connectivity approaches, highlighting their applications, advantages over traditional methods, and contributions to advancing neuroscientific knowledge.
The MCWG Outreach Council invites you to submit announcements or information about papers, conferences, presentations or other events or news related to brain and molecular connectivity as well as any positions available or job opportunities that you wish to publicize and share with the community!
Please submit any material for consideration by the final day of each month using this form – thank you!

The MCWG is made up of four international and multidisciplinary councils dedicated to promoting molecular connectivity research via dissemination of methods, results, collaboration, and resource sharing (e.g. datasets, tools) within the scientific community. We encourage the neuroscientific community to take an integrative perspective in study of the brain connectome, where various methods including MRI-based techniques, electrophysiological tools, and molecular imaging advance our understanding of the brain. Please find fundamental questions outlined here: “Brain connectomics: time for a molecular imaging perspective?”
Our website can be found here. We also invite you to join the MCWG!
[MCWG] Molecular Connectivity Newsletter: January 2025

Season’s Greetings, we would like to wish everyone a very happy and healthy new year!
MCOS returns this month and we hope you will join us on Friday, January 17, 2025, 22:00 15:00 CEST, 9:00 EDT with Mary C. Catanese, PhD (Department of Psychiatry, Massachusetts General Hospital, Harvard Medical School) who will discuss “Epigenetic alterations in white matter with age: impacts on structural connectivity and beyond”.
The registration link for the January MCOS can be found here!
We would also like to remind you that a preprint of an MCWG article is now available, further details below!
Title: Molecular connectivity studies in neurotransmission: a scoping review
Mario Severino, Débora Elisa Peretti, Marjorie Bardiau, Carlo Cavaliere, Matthieu Doyen, Gabriel Gonzalez-Escamilla, Tatiana Horowitz, Martin Nørgaard, Jhony Alejandro Mejia Perez, Matej Perovnik, Michael Rullmann, Dilara Steenken, Daniel Talmasov, Chunmeng Tang, Tommaso Volpi, Zhilei Xu, Alessandra Bertoldo, Vince D. Calhoun, Silvia Paola Caminiti, Xin Di, Christian Habeck, Sharna Jamadar, Daniela Perani, Arianna Sala, Vesna Sossi, Igor Yakushev, Joana B. Pereira, Mattia Veronese
1. We are pleased to announce the next MCOS talk featuring Mary C. Catanese, PhD!
Date: January 17th, 2025
Time: 22:00 AEST, 15:00 CET, 9:00 EST
Title: Epigenetic alterations in white matter with age: impacts on structural connectivity and beyond
Please join us for this 30 minute presentation to be followed by discussion (~25 minutes).
Please register here.
Abstract: Age related structural change in the healthy human brain has been associated with cognitive decline, however associated molecular alterations remain poorly understood. Positron emission tomography (PET) enables the in vivo investigation of molecular targets and can accelerate therapeutic development through their quantification. It has recently become possible to measure epigenetic machinery in the living human brain using the PET radiotracer [11C]Martinostat. This imaging agent targets a subset of class I histone deacetylases or HDACs. Epigenetic deacetylation of lysine residues of histone tails modifies chromatin in the regulation of gene expression. I am interested in HDACs as they are heritable and play critical roles in our response to the environment and are important in relation to cognitive processes including learning, memory and synaptic plasticity. Moreover, HDACs are implicated in psychiatric and neurodegenerative disorders. In previous work an age-related increase in [11C]Martinostat uptake was observed with age in healthy individuals and localized to the white matter. This increase in uptake correlated and co-localized with decreased structural integrity. I will discuss my goal to extend this work to study epigenetic alterations in relation to major depressive disorder by examining the potential association of epigenetic alterations (altered uptake) in relation to symptom related networks. I will also discuss using imaging-guided postmortem research programs initiated for the dual purpose of informing data interpretation and interrogating HDAC-related biology. My ultimate goal is to help better understand molecular dysregulation in psychiatric disorders.

Mary Catherine Catanese, PhD is Faculty Instructor in the Department of Psychiatry Department of Psychiatry, Massachusetts General Hospital, Harvard Medical School. She received her Ph.D. in Neuroscience and Behavior from the University of Massachusetts Amherst. Her work is focused on better understanding molecular dysregulation in psychiatric disorders using a multidimensional approach, integrating neuroscience, neuroimaging and clinical questions to address patient needs.
The MCOS promotes rigor in research and resource sharing. We aim to hold MCOS every third Friday of the month, subject to change due to speaker availability.
Please join us for February’s MCOS:
Please join us for our upcoming MCOS talks:
Please stay tuned for MCOS updates and reminders on social media!
2. We hope you will take a moment to read the MCWG authored preprint.
Molecular connectivity studies in neurotransmission: a scoping review


Purpose: Positron emission tomography (PET) and single photon emission computed tomography (SPECT) are essential molecular imaging tools for the in vivo investigation of neurotransmission. Traditionally, PET and SPECT images are analysed in a univariate manner, testing for changes in radiotracer binding in regions or voxels of interest independently of each other. Over the past decade, there has been an increasing interest in the so-called molecular connectivity approach that captures relationships of molecular imaging measures in different brain regions. Targeting these inter-regional interactions within a neuroreceptor system may allow to better understand complex brain functions. In this article, we provide a comprehensive review of molecular connectivity studies in the field of neurotransmission. We examine the expanding use of molecular connectivity approaches, highlighting their applications, advantages over traditional methods, and contributions to advancing neuroscientific knowledge.
The MCWG Outreach Council invites you to submit announcements or information about papers, conferences, presentations or other events or news related to brain and molecular connectivity as well as any positions available or job opportunities that you wish to publicize and share with the community!
Please submit any material for consideration by the final day of each month using this form – thank you!

The MCWG is made up of four international and multidisciplinary councils dedicated to promoting molecular connectivity research via dissemination of methods, results, collaboration, and resource sharing (e.g. datasets, tools) within the scientific community. We encourage the neuroscientific community to take an integrative perspective in study of the brain connectome, where various methods including MRI-based techniques, electrophysiological tools, and molecular imaging advance our understanding of the brain. Please find fundamental questions outlined here: “Brain connectomics: time for a molecular imaging perspective?”
Our website can be found here. We also invite you to join the MCWG!
[MCWG] Molecular Connectivity Newsletter: December 2024

Greetings to all!
We would like to wish everyone a very happy holiday season and we send our wishes for a happy and healthy new year!

Thank you to everyone who attended our MCOS in November with Hamish Deery who discussed “Metabolic connectivity in ageing”. The recording from his talk can be found here and the slides here.
As a note, MCOS will be taking a brief hiatus in December.
We hope you will join us for our next MCOS on Friday, January 17, 2025, 22:00 15:00 CEST, 9:00 EDT with Mary C. Catanese, PhD (Department of Psychiatry, Massachusetts General Hospital, Harvard medical school) who will discuss “Epigenetic alterations in white matter with age: impacts on structural connectivity and beyond”.
The registration link for the January MCOS can be found here!
1. We are pleased to announce the next MCOS talk featuring Mary C. Catanese, PhD!
Date: January 17th, 2025
Time: 22:00 AEST, 15:00 CET, 9:00 EST
Title: Epigenetic alterations in white matter with age: impacts on structural connectivity and beyond
Please join us for this 30 minute presentation to be followed by discussion (~25 minutes).
Please register here.
Abstract: Age related structural change in the healthy human brain has been associated with cognitive decline, however associated molecular alterations remain poorly understood. Positron emission tomography (PET) enables the in vivo investigation of molecular targets and can accelerate therapeutic development through their quantification. It has recently become possible to measure epigenetic machinery in the living human brain using the PET radiotracer [11C]Martinostat. This imaging agent targets a subset of class I histone deacetylases or HDACs. Epigenetic deacetylation of lysine residues of histone tails modifies chromatin in the regulation of gene expression. I am interested in HDACs as they are heritable and play critical roles in our response to the environment and are important in relation to cognitive processes including learning, memory and synaptic plasticity. Moreover, HDACs are implicated in psychiatric and neurodegenerative disorders. In previous work an age-related increase in [11C]Martinostat uptake was observed with age in healthy individuals and localized to the white matter. This increase in uptake correlated and co-localized with decreased structural integrity. I will discuss my goal to extend this work to study epigenetic alterations in relation to major depressive disorder by examining the potential association of epigenetic alterations (altered uptake) in relation to symptom related networks. I will also discuss using imaging-guided postmortem research programs initiated for the dual purpose of informing data interpretation and interrogating HDAC-related biology. My ultimate goal is to help better understand molecular dysregulation in psychiatric disorders.

Mary Catherine Catanese, PhD is Faculty Instructor in the Department of Psychiatry Department of Psychiatry, Massachusetts General Hospital, Harvard Medical School. She received her Ph.D. in Neuroscience and Behavior from the University of Massachusetts Amherst. Her work is focused on better understanding molecular dysregulation in psychiatric disorders using a multidimensional approach, integrating neuroscience, neuroimaging and clinical questions to address patient needs.
The MCOS promotes rigor in research and resource sharing. We aim to hold MCOS every third Friday of the month, subject to change due to speaker availability.
Please stay tuned for the announcement of the upcoming MCOS and for reminders on social media!
2. We are pleased to announce the preprint of an MCWG article.
Title: Molecular connectivity studies in neurotransmission: a scoping review
Authors: Mario Severino, Débora Elisa Peretti, Marjorie Bardiau, Carlo Cavaliere, Matthieu Doyen, Gabriel Gonzalez-Escamilla, Tatiana Horowitz, Martin Nørgaard, Jhony Alejandro Mejia Perez, Matej Perovnik, Michael Rullmann, Dilara Steenken, Daniel Talmasov, Chunmeng Tang, Tommaso Volpi, Zhilei Xu, Alessandra Bertoldo, Vince D. Calhoun, Silvia Paola Caminiti, Xin Di, Christian Habeck, Sharna Jamadar, Daniela Perani, Arianna Sala, Vesna Sossi, Igor Yakushev, Joana B. Pereira, Mattia Veronese
The MCWG Outreach Council invites you to submit announcements or information about papers, conferences, presentations or other events or news related to brain and molecular connectivity as well as any job opportunities that you wish to share with the community!
Please share for consideration by the final day of each month using this form.

The MCWG is made up of four international and multidisciplinary councils dedicated to promoting molecular connectivity research via dissemination of methods, results, collaboration, and resource sharing (e.g. datasets, tools) within the scientific community. We encourage the neuroscientific community to take an integrative perspective in study of the brain connectome, where various methods including MRI-based techniques, electrophysiological tools, and molecular imaging advance our understanding of the brain. Please find fundamental questions outlined here: “Brain connectomics: time for a molecular imaging perspective?”
Our website can be found here. We also invite you to join the MCWG!
[MCWG] Molecular Connectivity Newsletter: November 2024

Greetings!
Many thanks to all who participated in our Roundtable on Molecular Imaging of Brain Connectivity at EANM24 in Hamburg! We are grateful for the excellent feedback from all attendees and the many thought-provoking questions during the discussion!



Thank you to everyone who attended our MCOS in October with Igor Yakushev, MD, who discussed findings related to test-retest reproducibility of structural, functional and molecular estimates of brain connectivity at rest.
We hope you will join us for our next MCOS on Friday, November 22nd, 2024, please note a change in time as it will be held at 22:00 AEST, 14:00 CET, 8:00 EST with Hamish Deery (University of Monash, Australia) who will discuss “Metabolic connectivity in ageing”. As a note, MCOS will be taking a brief hiatus in December.
Please find details and the registration link for the November MCOS below!
We are pleased to announce the next MCOS talk featuring Hamish Deery!
Date: November 22nd, 2024
Time: 22:00 AEST, 14:00 CET, 8:00 EST
Title: Metabolic connectivity in ageing
Please join us for this 30 minute presentation to be followed by discussion (~25 minutes).
Please register here.
Abstract: Information transfer across the brain has a high energetic cost and requires efficient glucose metabolism. Here we use recently developed high temporal resolution functional positron emission tomography (fPET) to create a time course of glucose metabolism for individual subjects and assess the relationship between metabolic connectivity and cognitive function in ageing. The metabolic connectomes of 40 younger (mean 27.9 years; range 20-42) and 46 older (mean 75.8; 60-89) adults were characterised by high connectivity in the frontal, temporal, motor, parietal and medial cortices. Older age was associated with lower global integration of metabolic hub regions. In younger adults, a high proportion of glucose was used to support hubs in the frontal regions. Older adults used a higher proportion of a smaller energy budget to support mostly posterior hub regions. This metabolic network topology of older adults was associated with worse cognitive performance. We conclude that ageing is associated with a high glucose cost in hub regions and disrupted information transfer across the metabolic network. Our results highlight the fundamental role that metabolism plays in supporting information transfer in the brain and the unique insights that metabolic connectivity provides into the ageing brain.

Hamish Deery is a Ph.D. candidate in the School of Psychological Sciences and the Cognitive Neuroimaging Lab led by A/Professor Sharna Jamadar at Monash Biomedical Imaging, Monash University, in Melbourne, Australia. His research interests are in understanding brain function and cognition across the adult lifespan, including unique and potential shared pathways in ageing and lifestyle diseases. His work includes the use of simultaneous functional PET and MR to investigate the impact of insulin resistance on cerebral gluose metabolism and cognition in mid- and later-life.
The MCOS promotes rigor in research and resource sharing. We aim to hold MCOS every third Friday of the month, subject to change due to speaker availability.
Please note, MCOS will be taking a brief hiatus in December and will return in the new year, please join us on January 17th at 15:00 CET, 9:00 EST:
Mary Catherine Catanese, PhD (Department of Psychiatry, Massachusetts General Hospital, Harvard Medical School) will discuss Epigenetic alterations in white matter with age: impacts on structural connectivity and beyond
Please stay tuned for updates and reminders on social media!
In case you missed it, our educational course and symposium from OHBM2024 are now online!
Please find the educational course here and symposium here.

1. The Bini Lab in the Yale PET Center is happy to announce a job offering for a postdoctoral associate. The Bini Lab focuses on preclinical and clinical PET/CT and PET/MR imaging for applications in diabetes and obesity. The primary project examines changes in the dopaminergic system in the pancreas and brain during diabetes using PET/CT and MRI. Several additional on-going studies include PET/MR/CT imaging in liver, adipose tissue, and brain during obesity.
Responsibilities involve working with preclinical animal models, preclinical and clinical PET imaging, image analysis, kinetic modeling, data analysis, and results dissemination. The ideal candidate will have some prior hands-on experience in preclinical PET/CT or PET/MR imaging and kinetic modeling. A PhD in biomedical engineering, physics, or a related field is required. However, interested candidates with a strong computational background and interest in getting involved in medical imaging and preclinical models are also strongly encouraged to apply. Any wet lab experience for ex vivo tracer validation studies is a plus.
The successful candidate will – based on their research interest – have the option of participating in other PET studies as well, for example, PET connectivity, image-derived input functions and/or low-dose PET simulations.
More info:
https://medicine.yale.edu/profile/jason-bini
https://medicine.yale.edu/pet

2. Dr. Arianna Sala, is happy to announce a job offering for the support of up to two current graduate students/research assistants with a background in neuroscience, medicine, engineering and related fields, interested in applying for a PhD position (2025-2029) at the University of Liége, Belgium on a project related to either:
(1) Positron Emission Tomography (incl. molecular connectivity) in disorders of consciousness;
(2) multimodal PET/MR/EEG imaging (including all flavours of connectivity) in healthy volunteers.
If interested, please reach out via email: arianna.sala@uliege.be by November 17th. Please attach your CV and a brief cover letter discussing your interest.
Candidates who obtained a Master’s degree in 2024 (or expecting to graduate in 2025) and interested in a traineeship of up to 12 months with the Erasmus+ program (open to EU and non-EU candidates) are also welcome to apply.
The MCWG Outreach Council invites you to submit announcements or information about papers, conferences, presentations or other events or news related to brain and molecular connectivity as well as any job opportunities that you wish to share with the community!
Please share for consideration by the final day of each month using this form.

The MCWG is made up of four international and multidisciplinary councils dedicated to promoting molecular connectivity research via dissemination of methods, results, collaboration, and resource sharing (e.g. datasets, tools) within the scientific community. We encourage the neuroscientific community to take an integrative perspective in study of the brain connectome, where various methods including MRI-based techniques, electrophysiological tools, and molecular imaging advance our understanding of the brain. Please find fundamental questions outlined here: “Brain connectomics: time for a molecular imaging perspective?”
Our website can be found here. We also invite you to join the MCWG!
[MCWG] Molecular Connectivity Newsletter: October 2024

Hello!
We are excited to begin our 2024-2025 Molecular Connectivity Online Series, MCOS… again! We are very sorry for any convenience that may have arisen last month as we had to move the first talk of the season to October!
And so, without further ado, please join us on Friday October 18th, 2024, 15:00 CEST, 9:00 EDT for our MCOS with Igor Yakushev, MD, who will discuss findings related to test-retest reproducibility of structural, functional and molecular estimates of brain connectivity at rest.
Also, we are pleased to share that our educational course and symposium for OHBM2024 are now online! Please find the educational course here and symposium here.
Please find further details and registration link for MCOS below!
We are pleased to announce the next MCOS talk featuring Igor Yakushev, MD!
Date: October 18th, 2024
Time: 15:00 CET, 9:00 EST
Title: Test-retest reproducibility of structural and proxy estimates of brain connectivity at rest
Please join us for this 30 minute presentation to be followed by discussion (~25 minutes).
Please register here.
Abstract: While structural connectivity (SC) is indicative of actual anatomical connectivity and derived at a single subject level, proxy estimates of brain connectivity such as functional connectivity (FC) from functional magnetic resonance imaging (MRI), intersubject covariance of regional gray matter volume (GMVcov) from structural MRI, and intersubject covariance of regional 18F-fluorodeoxyglucose uptake (FDGcov) from positron emission tomography are derived from statistical dependencies between regional measurements. To understand, in how far these estimates are able to capture physiological and pathological changes in brain connectivity, knowledge on their reproducibility is essential. In this study, we determined reproducibility of group level SC, FC, GMVcov, and FDGcov in the same 55 healthy subjects at rest using a simultaneous PET/MRI acquisition protocol. Reproducibility was determined using Spearman’s correlation coefficient, coefficient of variation, and proportion of repeatedly (test and retest) present connections.

Igor Yakushev obtained his MD in 2010 from the University of Mainz, Germany. In 2012, after residency in Psychiatry and Neurology, he moved to Munich for residency in Nuclear Medicine at the Technical University of Munich (TUM), Germany. Since 2013 he has led the research group “Multimodal imaging of normal and pathological cognition”. Since 2015 he is the Head of Neuroimaging, since 2018 Senior Consultant in Nuclear Medicine at the Dept. of Nuclear Medicine, TUM. He is an Associate Faculty at the Munich Center for NeuroSciences “Brain and Mind”. Igor Yakushev has served in the Board of Directors of the Brain Imaging Council, Society of Nuclear Medicine and Molecular Imaging, as well as in the Neuroimaging Committee, European Association of Nuclear Medicine. His research is focused on mechanisms of brain connectivity and development of imaging-based biomarkers for neurodegenerative and neuro-oncological disorders.
You can find more about Igor here!
The MCOS promotes rigor in research and resource sharing. We aim to hold MCOS every third Friday of the month, subject to change due to speaker availability.
The following MCOS seminar will be held on Friday, November 22nd, 2024, and please note a change in time as it will be held at 22:00 AEST, 14:00 CET, 8:00 EST
Hamish Deery (University of Monash, Australia) will discuss “Metabolic connectivity in ageing”
Please stay tuned for reminders on social media!
We are excited to announce MCWG members will hold a Round Table at the 37th Annual Congress of the European Association of Nuclear Medicine (EANM), Hamburg, Germany October 19-23, 2024
Date: Monday October 21th, 2024
Time: 09:45 – 11:15 CET
Title: Round Table: Molecular Imaging of Brain Connectivity
Moderators: Silvia Morbelli (Turin, Italy) Igor Yakushev (Munich, Germany)
Arianna Sala (Liege, Belgium): Brain PET imaging 2.0: how can molecular imaging capture brain connectivity?
Alexander Drzezga (Cologne, Germany): Disease-specific patterns of brain dysconnectivity
Jorge Sepulcre (New Haven, United States of America): Establishing causal spreading of tau and amyloid with molecular imaging
More information can be found here.
Please visit MCWG members and colleagues who are presenting:
Silvia Paola Caminiti, Débora E. Peretti and Chunmeng Tang
Anna Lisa Martini, Giulia Carli, Silvia Paola Caminiti, Lorenzo Kiferle, Andrea Leo, Daniela Perani, Stelvio Sestini
Title: Brain metabolic connectivity and neurological symptoms persistence in Long-COVID
Presentation Number: OP-307 (TROP Session: Inflammation & Infection Committee: Top on Inflammation and Infection Imaging)
Time and location: Monday, October 21, 2024 8:00:00 AM – 9:30:00 AM, Hall Y10-Y12
Topic: This study delves into brain metabolic connectivity of SARS-CoV-2 survivors from acute-subacute to chronic phase aiming to shed light on the mechanisms underlying the persistence of neurological symptoms in long-COVID patients.
Débora E. Peretti, Sanne K. Meles, David Vállez García, Giulia Carli, Hans J. Van der Horn, Klaus L. Leenders, Remco J. Renken
Title: Open-source MATLAB code for the identification of spatial covariance patterns in neuroimaging data
Poster: EP-0764 (Session EP-51)
Date and time: The poster will be accessible throughout the conference at the e-poster area
Topic: An open-source SSM/PCA software easy to apply to difference radiotracers and disorders.
Chunmeng Tang, Michel Koole, Koen Van Laere, Philip Van Damme, Joke De Vocht
Title: Support Vector Machine Classification of 18F-FDG PET Scans in ALS Patients with and without C9orf72 Mutations versus Controls.
Presentation Number: OP-857 (Session: 1805 Cutting Edge Science Track – TROP Session: Physics Committee: Data Analysis: Neuro & Cardio)
Time and location: Wednesday, 23/10, 9:45:00 AM – 11:15:00 AM, Hall Y4-Y9
Topic: The classification between C9-ALS and sALS achieved accuracies higher than 70% for both classes, showing that the pattern of brain involvement in C9-ALS considerably differs from sALS.
The MCWG Outreach Council invites you to submit information about papers, conferences, presentations or other events or news related to brain and molecular connectivity as well as any job opportunities that you wish to share with the community!
Please share for consideration by the final day of each month using this form.

The MCWG is made up of four international and multidisciplinary councils dedicated to promoting molecular connectivity research via dissemination of methods, results, collaboration, and resource sharing (e.g. datasets, tools) within the scientific community. We encourage the neuroscientific community to take an integrative perspective in study of the brain connectome, where various methods including MRI-based techniques, electrophysiological tools, and molecular imaging advance our understanding of the brain. Please find fundamental questions outlined here: “Brain connectomics: time for a molecular imaging perspective?”
Our website can be found here. We also invite you to join the MCWG!
[MCWG] Molecular Connectivity Newsletter: September 2024

Guess who’s back!!!
We hope that you have had a safe and restful summer! We are excited to resume our monthly newsletter for the academic year! We are also resuming our Molecular Connectivity Online Series, MCOS!
Thank you to everyone who attended the June MCOS with Tommaso Volpi, PhD, the recording from his talk can be found here and slides here.
Please stay tuned for further announcements as well as links to recordings from MCOS talks and last year’s Symposium coming soon!
Please join us on Friday September 20th, 2024, 15:00 CEST, 9:00 EDT for our MCOS with Igor Yakushev, MD, who will discuss findings related to test-retest reproducibility of structural, functional and molecular estimates of brain connectivity at rest.
Please find further details and registration information below!
We are pleased to announce the next MCOS talk featuring Igor Yakushev, PhD!
Date: September 20th, 2024
Time: 15:00 CET, 9:00 EST
Title: Test-retest reproducibility of structural and proxy estimates of brain connectivity at rest
Please join us for this 30 minute presentation to be followed by discussion (~25 minutes).
Please register here.
Abstract: While structural connectivity (SC) is indicative of actual anatomical connectivity and derived at a single subject level, proxy estimates of brain connectivity such as functional connectivity (FC) from functional magnetic resonance imaging (MRI), intersubject covariance of regional gray matter volume (GMVcov) from structural MRI, and intersubject covariance of regional 18F-fluorodeoxyglucose uptake (FDGcov) from positron emission tomography are derived from statistical dependencies between regional measurements. To understand, in how far these estimates are able to capture physiological and pathological changes in brain connectivity, knowledge on their reproducibility is essential. In this study, we determined reproducibility of group level SC, FC, GMVcov, and FDGcov in the same 55 healthy subjects at rest using a simultaneous PET/MRI acquisition protocol. Reproducibility was determined using Spearman’s correlation coefficient, coefficient of variation, and proportion of repeatedly (test and retest) present connections.

Igor Yakushev obtained his MD in 2010 from the University of Mainz, Germany. In 2012, after residency in Psychiatry and Neurology, he moved to Munich for residency in Nuclear Medicine at the Technical University of Munich (TUM), Germany. Since 2013 he has led the research group “Multimodal imaging of normal and pathological cognition”. Since 2015 he is the Head of Neuroimaging, since 2018 Senior Consultant in Nuclear Medicine at the Dept. of Nuclear Medicine, TUM. He is an Associate Faculty at the Munich Center for NeuroSciences “Brain and Mind”. Igor Yakushev has served in the Board of Directors of the Brain Imaging Council, Society of Nuclear Medicine and Molecular Imaging, as well as in the Neuroimaging Committee, European Association of Nuclear Medicine. His research is focused on mechanisms of brain connectivity and development of imaging-based biomarkers for neurodegenerative and neuro-oncological disorders.
You can find more about Igor here!
The MCOS promotes rigor in research and resource sharing. We aim to hold MCOS every third Friday of the month, subject to change due to speaker availability.
The following MCOS seminar will be held on Friday, October 18th, 2024, 15:00 CEST, 9:00 EDT
Mary C. Catanese, PhD (Department of Psychiatry, Massachusetts General Hospital, Harvard Medical School, USA) will discuss “Epigenetic alterations in white matter with age: impacts on structural connectivity and beyond”
The subsequent MCOS will be held on Friday, November 22nd, 2024, and please note a change in time as it will be held at 22:00 AEST, 14:00 CET, 8:00 EST
Hamish Deery (University of Monash, Australia) will discuss “Metabolic connectivity in ageing”
Please visit us on social media and stay tuned for further details!
The MCWG Outreach Council invites you to submit information about papers, conferences, presentations or other events or news related to brain and molecular connectivity as well as any job opportunities that you wish to share with the community!
Please share for consideration by the final day of each month using this form.

The MCWG is made up of four international and multidisciplinary councils dedicated to promoting molecular connectivity research via dissemination of methods, results, collaboration, and resource sharing (e.g. datasets, tools) within the scientific community. We encourage the neuroscientific community to take an integrative perspective in study of the brain connectome, where various methods including MRI-based techniques, electrophysiological tools, and molecular imaging advance our understanding of the brain. Please find fundamental questions outlined here: “Brain connectomics: time for a molecular imaging perspective?”
Our website can be found here. We also invite you to join the MCWG!
[MCWG] Molecular Connectivity Newsletter: June 2024

Hello!
Many thanks to all who participated in our Molecular Connectivity Symposium, “What is brain connectivity?” ! We wish to thank our generous sponsors, DFG, Bruker (PMOD), Curium and Technical University of Munich!

After the excellent talks and discussions at the Symposium, we have set a foundation for a forthcoming nomenclature consensus paper, stay tuned!
Also after the success of the symposium in Munich, we continue our discussions online with the Molecular Connectivity Online Series (MCOS)! Thanks to all who attended the May MCOS with Joana Pereira, PhD!
Please join us for our final MCOS of the season in June, featuring Tommaso Volpi, PhD! Please find details and registration information below!
We are also sharing details about MCWG-related events at the upcoming OHBM Meeting, with information about talks as well as posters from within and outside of MCWG in areas related to molecular questions and brain connectivity. We hope to see you at MCOS and at OHBM, in Seoul, Korea!
As a final note, the newsletter will be quiet for the next few months for a summer hiatus and will return in the fall! Please stay tuned for announcements as well as links to recordings from the Symposium coming in September!
We hope everyone has a safe and restful summer!
We are pleased to announce the next MCOS talk featuring Tommaso Volpi, PhD!
Date: June 14th, 2024
Time: 15:00 CET, 9:00 EST
Title: Molecular connectivity & dynamic PET: comparing time series and subject series approaches
Please join us for this 30 minute presentation to be followed by discussion (~25 minutes).
Please register here.
Dynamic PET acquisitions allow individuals to obtain individual estimates of molecular connectivity (MC) from time series data – similarly to fMRI functional connectivity. In this talk, focusing on [18F]FDG as a representative tracer, I will discuss
Much of this content is included in our 2023 JCBFM paper.

Dr. Tommaso Volpi is currently a Postdoctoral Associate at the Department of Radiology at Yale University, under the supervision of Prof. Richard Carson. He obtained his Ph.D. in 2023 from the University of Padova, Italy, with a thesis on the complex coupling between [18F]FDG PET measures of brain glucose metabolism and fMRI proxies of spontaneous activity, including the comparison of metabolic and functional connectivity. His main current projects concern methods for PET kinetic modeling and noninvasive input function estimation to obtain physiologically informative parameters, and the integration of PET- and MRI-derived features to understand brain function and connectivity in health and disease (Parkinson’s disease, epilepsy, gliomas). He is a member of the Validation Council of the MCWG.
You can find more about Tommaso here!
The MCOS promotes rigor in research and resource sharing. We aim to hold MCOS every third Friday of the month, but is subject to change due to speaker availability.
The next MCOS seminar will be held in September, 2024!
Please watch our social media and stay tuned for further details!
1. OHBM 2024 will feature an educational course organized by members of the MCWG:
Molecular Imaging: a Hands-on Tutorial based on Open Access Datasets
Date: June 23 2024
Time: 1:30 PM – 5:30 PM
Room: Grand Ballroom 105

Organized by Arianna Sala and Igor Yakushev, the educational course will feature presentations from Arianna Sala, Alessandra Bertoldo, Martin Nørgaard, Xin Di and Granville Matheson. This Educational Course aims to provide a basic introduction to molecular imaging, covering its basic principles and main tools, illustrating similarities and critical differences in the main molecular measures commonly encountered in open access/easily accessible molecular datasets. Different hands-on, interactive sessions will allow the attendees to get first-hand experience on data access, processing and modelling of different types of molecular imaging data. The target audience for this educational course are researchers in the field of neuroimaging of all levels of expertise, wishing to expand their knowledge.
Presentations:
For further information about the Educational course, please see here.
2. OHBM 2024 will also feature a symposium organized by members of the MCWG:
Brain connectivity: Ready for clinical applications?
An overview of current challenges and caveats
Date: June 24 2024
Time: 8:00 AM – 9:15 AM
Room: Grand Ballroom 103

Organized by Débora Peretti and Arianna Sala, the symposium is entitled “Brain connectivity: ready for clinical application? An overview of current challenges and caveats” and will feature presentations from Simon Eickhoff, Christian Habeck, Alessandro Gozzi, and Débora Peretti. In this symposium we discuss some of the most pressing methodological considerations that still need to be addressed by neuroscientists in order to obtain robust connectivity markers for inference and prediction. Both statistical and biological considerations will be covered, on the grounds that not only prediction of phenotypes from activation/connectivity data, but also understanding the mechanisms underlying inter-individual connectivity differences, are necessary to obtain robust, generalizable topographic substrates. Finally, an overview of the translation of connectivity metrics into clinical practice will be presented.
Presentations:
For further information about the Symposium, please see here.
3. MCWG Member Posters at OHBM
Débora Peretti, Cecilia Boccalini, Max Scheffler, Cristelle Rodriguez, Marie Montandon, Sven Haller, Panteleimon Giannakopoulos, Giovanni B. Frisoni, Valentina Garibotto
Title: Brain Connectivity Measured Through Spatial Covariance as a Biomarker for Neurodegeneration in AD
Poster number: 252
Date and time: June 24th, 01:15 – 03:15 PM, and June 25th 02:00 – 04:00 PM
Topic: Spatial covariance maps provide a strong and specific marker for AD, outperforming conventional metrics to assess neurodegeneration.
Amritha Harikumar, Maria Misiura, Daniel Amen, David Keator, Vince Calhoun
Title: SPECT brain networks and their functional connectivity in schizophrenia patients vs controls.
Poster number: 661
Date and time: June 24th, 2024: 01:15 PM – 03:15 PM and June 25th, 2024: 02:00 PM – 04:00 PM
Topic: Here we utilize SPECT data to compare group differences in patients with schizophrenia and healthy controls using fully automated, spatially constrained ICA (i.e., the Neuromark pipeline). We evaluate both the spatial regions as well as the whole brain SPECT connectome (assessed as covariation among subjects) to evaluate the neuroimaging links to schizophrenia
Giulia Vallini, Giorgia Baron, Giulia Pagnin, John J. Lee, Andrei G. Vlassenko, Manu S. Goyal, Diego Cecchin, Maurizio Corbetta, Alessandra Bertoldo
Title: Whole-brain metabolic-effective coupling in healthy controls and its disruption in gliomas.
Poster number: 2492
Date and time: June 26th, 2024: 12:45 PM – 14:45 PM and June 27th, 2024: 12:45 PM – 14:45 PM
Topic: In our study, a dual metabolic-functional association emerges: one at the local level (glucose uptake-nodal signal directionality) and one at the network level (metabolic connectivity-partial covariance). Furthermore, our study reveals how these two distinct decouplings can differentiate glioma patients based on lesion location – with local disruptions observed only for temporo-parietal lesions and network alterations for frontal lesions – providing new insights into the pathophysiology and metabolic-kinetic linkage of glioma
4. Molecular Connectivity Related Posters at OHBM
Penghui Du, Sean Coursey, Ting Xu, Hsiao-Ying Wey, Jonathan Polimeni, Quanying Liu, Jingyuan Chen
Title: Human Cerebral Cortex Organization Estimated by Functional PET-FDG “Metabolic Connectivity”
Poster number: 1496
Date and time: June 26, 2024: 01:45 PM – 03:45 PM and June 27, 2024: 01:45 PM – 03:45 PM
Topic: In this study, we applied a connectivity gradient-based analytical scheme on a resting-state simultaneous fPET-fMRI dataset, aiming to characterize the detailed cortical organization of fPET-derived MC and understand how it differs from the fMRI-derived functional network structures.
Fernando Bravo, Sabrina Haas, Tudor Ionescu, Irene Gonzalez-Menendez, Leticia Quintanilla-Martinez, Gina Dunkel, Laura Kuebler, Bettina Weigelin, Gerald Reischl, Bernd Pichler, Kristina Herfert
Title: Simultaneous fPET/fMRI unveils nigrostriatal pathway mechanisms during optogenetic stimulation.
Poster number: 2397
Date and time: June 26, 2024: 01:45 PM – 03:45 PM and June 27, 2024: 01:45 PM – 03:45 PM
Topic: In this study, the use of simultaneous [18F]FDG-fPET/BOLD-fMRI allowed us to demonstrate excitation-inhibition mechanisms of the nigrostriatal pathway during optogenetic stimulation, which are concealed in isolated fMRI readouts.
We invite you to please send information about events related to brain and molecular connectivity or news or job opportunities that you wish to share with the community for consideration by the final day of each month using this form.
The Functional Neuroimaging Laboratory has a postdoctoral position (up to 6 years, internationally competitive salary) aimed at understanding the biological basis of brain connectivity by implementing multimodal optical imaging and functional neuroimaging in the mouse – maybe of interest for member PhD students: https://www.fens.org/careers/job-market/job/118960
This position is part of a newly funded research program that is launching to understand how brain-wide networks respond to internal and external modulation. The lab (www.fnimg.iit.it) is a supportive and collaborative environment, and this project is part of a larger program with opportunities for future collaboration and growth. The lab expects to have multiple new openings soon (including 1 data analyst, 3 postdocs and 1 PhD student) in the coming weeks, so please stay tuned.
The MCWG Outreach Council invites you to submit information about papers, conferences, presentations or other events or news related to brain and molecular connectivity or job opportunities that you wish to share with the community for consideration by the final day of each month using this form.

The MCWG is made up of four international and multidisciplinary councils dedicated to promoting molecular connectivity research via dissemination of methods, results, collaboration, and resource sharing (e.g. datasets, tools) within the scientific community. We encourage the neuroscientific community to take an integrative perspective in study of the brain connectome, where various methods including MRI-based techniques, electrophysiological tools, and molecular imaging advance our understanding of the brain. Please find fundamental questions outlined here: “Brain connectomics: time for a molecular imaging perspective?”
Our website can be found here. We also invite you to join the MCWG!
[MCWG] Molecular Connectivity Newsletter: May 2024

Hello!
We wish to remind everyone that registration for the Molecular Connectivity Symposium, “What is brain connectivity?” is still open for online participation with ten spots available! The registration link can be found here, with more information below! We wish to ensure that everyone who is interested in attending online has the opportunity to attend and look forward to “seeing” you there! Also, please join us for our May MCOS with Dr. Joana Pereira, on Friday, May 17th, 15:00 CET, 9:00 EST, further details and registration link below!
1. The MCWG second Symposium “What is brain connectivity?” is this Friday May 3rd!
We hope you will join us!
Please note, there are 10 available spots remaining for online participation!
Registration: Registration for this symposium is free! Secure your spot by visiting here.
The symposium aims to better delineate macroscale brain connectivity across fMRI, DWI, EEG, and molecular connectivity. The event will provide a foundation for follow-up discussions about standardized nomenclature for molecular connectivity.
🌐 Please join Us!
Date: May 3, 2024
Time: 9:00 am – 1:00 pm (GMT+2, time converter: here)
We are also offering Special Access for Molecular Connectivity Members continuing through May the 4th: “Molecular Imaging of Brain Connectivity: towards standardized nomenclature” reserved for MCWG members and experts.
This event is organized with the support of DFG, Bruker (PMOD), Curium and Technical University of Munich.
2. We are pleased to announce the next MCOS talk featuring Joana Pereira, PhD!
Date: May 17th, 2024
Time: 15:00 CET, 9:00 EST
Title: Individual PET connectomes capture disease progression and cognitive decline in Alzheimer’s disease
Please join us for this 30 minute presentation to be followed by discussion (~25 minutes).
Please register here.
Identifying unique differences is crucial for enhancing personalized medicine strategies for Alzheimer’s disease (AD). In this work, we present a framework to create individual molecular connectomes by using longitudinal data from tau and amyloid positron emission tomography scans. We show that these personalized molecular connectomes can pinpoint specific individuals and track disease progression throughout different AD stages as well as over time.

I have a background in neuroimaging in aging and neurodegenerative diseases, mainly Alzheimer’s disease (AD). In my group we are working with different brain imaging modalities, CSF and plasma biomarkers to understand the sequence of pathological events that lead to AD. We have developed different methods, including an open-access software for the analysis of brain connectivity using graph theory, multilayer network approaches and deep learning called BRAPH.
The MCOS promotes rigor in research and resource sharing. We aim to hold MCOS every third Friday of the month, but is subject to change due to speaker availability.
Our next seminar will be held on Friday June 14th, 2024 at 15:00 CET, 9:00am EST featuring Tommaso Volpi, PhD who will discuss Molecular connectivity & dynamic PET: comparing time series and subject series approaches.
Please stay tuned for further details in next month’s newsletter!
Reminder, OHBM 2024 will feature a symposium organized by members of the MCWG:
Brain connectivity: Ready for clinical applications?
Date: June 24 2024
Time: 8:00 AM – 9:15 AM

Organized by Débora Peretti and Arianna Sala, the symposium is entitled “Brain connectivity: ready for clinical application? An overview of current challenges and caveats” and will feature presentations from Simon Eickhoff, Christian Habeck, Alessandro Gozzi, and Débora Peretti. In this symposium we discuss some of the most pressing methodological considerations that still need to be addressed by neuroscientists in order to obtain robust connectivity markers for inference and prediction. Both statistical and biological considerations will be covered, on the grounds that not only prediction of phenotypes from activation/connectivity data, but also understanding the mechanisms underlying inter-individual connectivity differences, are necessary to obtain robust, generalizable topographic substrates. Finally, an overview of the translation of connectivity metrics into clinical practice will be presented.
We invite you to please send information about events related to brain and molecular connectivity or news or job opportunities that you wish to share with the community for consideration by the final day of each month using this form.
We will feature a member of the MCWG and include a brief Q&A, stay tuned for next month’s feature!

The MCWG is made up of four international and multidisciplinary councils dedicated to promoting molecular connectivity research via dissemination of methods, results, collaboration, and resource sharing (e.g. datasets, tools) within the scientific community. We encourage the neuroscientific community to take an integrative perspective in study of the brain connectome, where various methods including MRI-based techniques, electrophysiological tools, and molecular imaging advance our understanding of the brain. Please find fundamental questions outlined here: “Brain connectomics: time for a molecular imaging perspective?”
Our website can be found here. We also invite you to join the MCWG!
[MCWG] Molecular Connectivity Newsletter: April 2024

Greetings!
Thanks to everyone in attendance at our MCOS in March with Vince Calhoun, PhD! Please find the recording here and the slides here. We welcome any feedback and hope to see you at our April MCOS seminar with Simon Eickhoff, PhD, next Friday 19 April! Time and registration information is below!
In this newsletter:
We are pleased to announce the fourth MCOS talk featuring Simon Eickhoff, PhD!
Date: April 19th, 2024
Time: 15:00 CET, 9:00 EST
Title: The many faces of brain connectivity
Please join us for this 30 minute presentation to be followed by discussion (~25 minutes).
Please register here.
Regional segregation and long-range integration are the two fundamental principles of brain organization. As connectivity patterns play a major role in both of these, assessing the the connectome has emerged as a key avenue towards a better understanding of the brain. Key to this presentation is the notion that there is no such thing as “the” connectivity between two brain regions, but rather different concepts, facets and assessment of brain connectivity may provide corroborating, complementary or conflicting evidence. Therefore, a better understanding of the relationship between connectivity-types and their integration to a more holistic view of long-range interactions should be critical to avoid an undue reliance on individual approaches that may actually hinder progress in neurobiological understanding.

Simon Eickhoff is a full professor and chair of the Institute for Systems Neuroscience at the Heinrich-Heine University in Düsseldorf and the director of the Institute of Neuroscience and Medicine (INM-7, Brain and Behavior) at the Forschungszentrum Jülich. He is furthermore a visiting professor at the Chinese Academy of Science Institute of Automation. Working at the interface between neuroanatomy, data-science and brain medicine, he aims to obtain a more detailed characterization of the organization of the human brain and its inter-individual variability in order to better understand its changes in advanced age as well as neurological and psychiatric disorders. This goal is pursued by the development and application of novel analysis tools and approaches for large-scale, multi-modal analysis of brain structure, function and connectivity as well as by machine-learning for single subject prediction of cognitive and socio-affective traits and ultimately precision medicine.
The MCOS promotes rigor in research and resource sharing. We aim to hold MCOS every third Friday of the month, but is subject to change due to speaker availability.
Our next seminar will be held on Friday May 17th, 2024 at 15:00 CET, 9:00am EST featuring Joana Pereira, PhD who will discuss her work on brain connectivity in Alzheimer’s Disease.
Stay tuned for details in next month’s newsletter!
1. MCWG symposium
We are pleased to remind you that the second MCWG symposium:
“What is Brain Connectivity?”
will be held in Munich on 3 May (9:00 am – 1:00 pm CEST).
Registration for this hybrid event is free but only a few places are left, so don’t miss out on this opportunity !
🚨 Important information : Registration for in-person attendance will close on April 14th.
You can secure your spot by registering here.
During this event, international speakers will delve into the fascinating world of brain connectivity. Our program promises enlightening discussions and valuable insights from experts in the fields of fMRI, DWI, EEG, and molecular imaging.
This symposium will be followed by Sessions reserved to MCWG members and experts dedicated to “Molecular Imaging of Brain Connectivity: towards standardized nomenclature“ (📅 May 3-4).
🌐 Programme and info: tinyurl.com/MCWGMunich2024
We look forward to welcoming you to our symposium and exploring the exciting world of brain connectivity together!
2. We are very pleased to announce that OHBM 2024 will feature a symposium organized by members of the MCWG!
Date: June 24 2024
Time: 8:00 AM – 9:15 AM
Title: Brain connectivity: Ready for clinical applications? An overview of current challenges and caveats

Organized by Débora Peretti and Arianna Sala, the symposium is entitled “Brain connectivity: ready for clinical application? An overview of current challenges and caveats” and will feature presentations from Simon Eickhoff, Christian Habeck, Alessandro Gozzi, and Débora Peretti. In this symposium we discuss some of the most pressing methodological considerations that still need to be addressed by neuroscientists in order to obtain robust connectivity markers for inference and prediction. Both statistical and biological considerations will be covered, on the grounds that not only prediction of phenotypes from activation/connectivity data, but also understanding the mechanisms underlying inter-individual connectivity differences, are necessary to obtain robust, generalizable topographic substrates. Finally, an overview of the translation of connectivity metrics into clinical practice will be presented.
We invite you to please send information about events related to brain and molecular connectivity or news or job opportunities that you wish to share with the community for consideration by the final day of each month using this form.
1. Job Opening at the University of Geneva
Would you like to study different pathological mechanisms underlying Alzheimer’s disease and its synergistic effect leading to clinical symptoms and progression?
The Neuroimaging and Innovative Molecular Tracers Lab at the University of Geneva, lead by Prof. Valentina Garibotto, is currently looking for an enthusiastic PhD candidate interested in learning about Positron Emission Tomography imaging and the risk factors leading to Alzheimer’s disease.
Apply at: https://jobs.unige.ch/www/wd_portal.show_job?p_web_site_id=1&p_web_page_id=64498
Please submit any news, including publications you would like highlighted or job listings related to brain connectivity and molecular connectivity that you would like to share with the community for consideration no later than the final day of each month using this form.
Stay tuned for upcoming features!

The MCWG is made up of four international and multidisciplinary councils dedicated to promoting molecular connectivity research via dissemination of methods, results, collaboration, and resource sharing (e.g. datasets, tools) within the scientific community. We encourage the neuroscientific community to take an integrative perspective in study of the brain connectome, where various methods including MRI-based techniques, electrophysiological tools, and molecular imaging advance our understanding of the brain. Please find fundamental questions outlined here: “Brain connectomics: time for a molecular imaging perspective?”
Our website can be found here. We also invite you to join the MCWG!
[MCWG] Molecular Connectivity Newsletter: March 2024

Hello!
We hope all is well! We wish to thank those who were able to attend our MCOS in February with Christian Habeck, PhD! Please find the recording here and the slides here. We welcome any feedback you may have and hope to see you at our March MCOS seminar. Please note, this month’s MCOS will be held earlier than usual, on March 15th, please find further details below!
1. We are pleased to inform you that the second Molecular Connectivity Working Group Symposium is set for May 3-4 in Munich, Germany
🌐 Join Us at the Molecular Connectivity Symposium in Munich!
📅 Date: May 3, 2024
🕒 Time: 9:00 am – 1:00 pm
📰 Programme
The Molecular Connectivity Working Group is delighted to unveil its second Symposium with the title “What is brain connectivity?”. The symposium aims to identify a common denominator in the definition of macroscale brain connectivity across fMRI, DWI, EEG, and molecular connectivity. The event is supposed to provide a baseline for follow-up debates on standardized nomenclature for molecular connectivity.
Key Highlights:
Registration:
📝 Registration for this symposium is now open and free! Secure your spot by visiting here. Don’t miss out on this opportunity to be a part of a groundbreaking exploration into molecular imaging and brain connectivity.
Exclusive Access for Molecular Connectivity Members:
The symposium is part of a larger event continuing till May the 4th titled “Molecular Imaging of Brain Connectivity: towards standardized nomenclature” exclusively reserved for MCWG members and experts, please join MCWG for access!
The event is organized with the support of DFG, Bruker, Curium and Technical University of Munich.
📍 See you in Munich!
2. We are also very pleased to announce the third MCOS talk featuring Vince Calhoun, PhD!
Date: March 15th, 2024 March 22nd, 2024
Time: 15:00 CET, 09:00 EST 10:00 EST
Title: NeuroMark PET: Towards a fully automated PET ICA pipeline
Please join us for this 30 minute presentation to be followed by discussion (~25 minutes).
Please register here.
Abstract: Anatomy based atlases are often used to summarize positron emission tomography (PET) data. However, it is well known that functional boundaries do not correspond well to anatomic boundaries. In addition, anatomic atlases do not capture variation among individuals and may average together voxels which are functionally distinct. In contrast, independent component analysis (ICA) provides informative data-driven output which also allows for overlap across anatomic regions. However, the output of blind ICA, without any a priori information, can be challenging to compare across studies and requires manual selection of components of interest. Here we propose the use of a spatially constrained ICA approach, called NeuroMark PET, leveraging a priori PET template derived from ICAs which replicated across independent PET datasets. We use this to generate a radioligand specific template which is then used to automatically generate component maps (i.e., covarying networks) and subject specific output using spatially constrained ICA. We also show that the ICA component maps are invariant to standard update value ratio (SUVR) scaling, allowing easy rescaling of the component loadings as desired. The proposed Neuromark PET approach effectively captures biologically meaningful subject-specific features that are comparable across different individuals. We demonstrate by comparing group differences in PET networks as well as age-related changes. This approach also allows for comparison across modalities or radioligands. We show an example of this by using a widely used functional MRI NeuroMark template as spatial priors for PET data, revealing PET specific modularity in fMRI intrinsic networks. In sum, we describe a PET NeuroMark template and pipeline and suggest that it represents a powerful resource for the research community and for analyses with large multimodal datasets.
Dr. Calhoun is founding director of the tri-institutional Center for Translational Research in Neuroimaging and Data Science (TReNDS) where he holds appointments at Georgia State, Georgia Tech and Emory. He is the author of more than 1000 full journal articles. His work includes the development of flexible methods to analyze neuroimaging data including blind source separation, deep learning, multimodal fusion and genomics, neuroinformatics tools. Dr. Calhoun is a fellow of the Institute of Electrical and Electronic Engineers, The American Association for the Advancement of Science, The American Institute of Biomedical and Medical Engineers, The American College of Neuropsychopharmacology, The Organization for Human Brain Mapping (OHBM) and the International Society of Magnetic Resonance in Medicine. He currently serves on the IEEE BISP Technical Committee and is also a member of IEEE Data Science Initiative Steering Committee as well as the IEEE Brain Technical Committee.
The MCOS promotes rigor in research and resource sharing. We aim to hold MCOS every third Friday of the month, but is subject to change due to speaker availability.
Our next seminar will be held on April 19th, 2024 at 15:00 CET, 9:00am EST featuring Simon Eickhof, PhD who will discuss The many faces of brain connectivity.
Stay tuned for details in next month’s newsletter!
We are pleased to announce that OHBM 2024 will feature a symposium organized by members of the MCWG!

Organized by Débora Peretti and Arianna Sala, the symposium is entitled “Brain connectivity: ready for clinical application? An overview of current challenges and caveats” and will feature presentations from Simon Eickhoff, Christian Habeck, Alessandro Gozzi, and Débora Peretti. In this symposium we discuss some of the most pressing methodological considerations that still need to be addressed by neuroscientists in order to obtain robust connectivity markers for inference and prediction. Both statistical and biological considerations will be covered, on the grounds that not only prediction of phenotypes from activation/connectivity data, but also understanding the mechanisms underlying inter-individual connectivity differences, are necessary to obtain robust, generalizable topographic substrates. Finally, an overview of the translation of connectivity metrics into clinical practice will be presented. Date and time to be announced.
Members of the MCWG are co-organizing the 12th European Conference on Clinical Neuroimaging in March in Leipzig, featuring invited talks on molecular imaging, total-body PET, imaging in psychiatry and ATN concept in Alzheimer’s disease. Register now.
Please send us information about events related to brain connectivity and molecular connectivity that you wish to share with the community for consideration no later than the final day of each month using this form.
Please submit any news, including publications you would like highlighted or job listings related to brain connectivity and molecular connectivity that you would like to share with the community for consideration no later than the final day of each month using this form.
Each month, we will feature a member of the MCWG and inclide a brief Q&A!
This month we highlight Dr. Sharna Jamadar, member of the MCWG Steering Committee.
Dr. Sharna Jamadar is Associate Professor (Research) and NHMRC Emerging Leader Fellow at Monash Biomedical Imaging and Turner Institute for Brain and Mental Health.Sharna’s research is focused on understanding how our life experiences change our brains, and how this may confer resilience to the ageing process. She is particularly interested in studying the neural bases of cognitive control, and has explored the consequences of disruption of cognitive control in a number of conditions, including healthy ageing, psychosis (schizophrenia, bipolar disorder) and substance abuse. She also has an emerging interest in the neuroscience of parenthood and is supervising two student projects in this theme. Sharna is an expert in multimodal neuroimaging, and uses a number of imaging techniques to understand cognitive control, including positron emission tomography (PET), functional magnetic resonance imaging (fMRI), electroencephalography (EEG) and oculomotor measures (eye-tracking). Sharna’s team leads the development of simultaneous functional MRI-PET imaging in humans at the facility. Sharna’s team has developed novel MRI-PET measures that provide high resolution mapping of the function, structure, and metabolic efficiency of the brain. For the first time, these new techniques allow researchers to study task-related changes in brain function and metabolism with a temporal resolution below 20sec. This work will have substantial implications for our understanding of how the brain dynamically uses energy; and Sharna will use these new methods to understand how the metabolic efficiency of the brain changes across the lifespan.
Dr. Sharna Jamadar has graciously responded to our feature questionnaire!
What sparked your interest in molecular imaging or led you to focus on research in molecular imaging?
I trained as a cognitive neuroscientist in EEG and fMRI, and at the time that I was doing my training (2005-2010), I (somewhat naively) thought of PET as a functional neuroimaging tool that had been superseded by fMRI. In terms of functional brain mapping and brain-cognition relationships, the vast majority of the work I was reading, and doing myself, was focused on electrophysiology and haemodynamics. From the beginning though, I always had a healthy dose of scepticism about what we can and can’t do with fMRI – you may remember a rather famous paper by Logothetis with that exact title.
In 2015 I returned to work after a year’s maternity leave, and this was a time of transition both personally and professionally. At that time, we were installing a new PET/MR scanner at our institute, and I was really excited about the possibilities this new technology was offering. In particular, I was intrigued by how FDG-PET can be considered a more direct and quantitative measure of neuronal function than fMRI. I was moving my research into the area of cognitive ageing, and this was exciting to me, as there are lots of challenges to using fMRI in older adults, particularly if we want to compare them to younger adults, which I did. I was particularly inspired by the Villien et al. 2014 paper in NeuroImage, with the proposed method for functional PET (fPET) for brain mapping. We noticed that their approach didn’t provide simultaneous task contrast for both fMRI and fPET, so our first pilot study was a simple replication and extension of that work to allow this. I’d never used PET before, let alone fPET, and this was a really fun time playing in the lab and figuring out this new method for measuring brain function. It was a great time – we tried lots of new things in the scanner, as well as with the data, collaborating across radiography, nuclear medicine technology, engineering, maths, and physics – with me the lowly neuroscientist trying to figure out what it all means!
As a follow-up to this work, I quickly shifted to considering how we might use PET to measure connectivity in the brain. The human brain mapping community had been really focused on resting connectivity for some time, and I’d done a bit of work in this area too. We’d just published a paper showing how the fMRI connectome is confounded by individual differences in haemoglobin in an older population, and so the idea that we might be able to get a less confounded measure of connectivity was really appealing. Of course, I was “scooped” back in the 1980s by Barry Horwitz and colleagues, as well as the greater PET community in the subsequent years. However, after reading the literature, I realised that the definition of connectivity used in PET was different to how I’d use it if I was looking at an EEG or fMRI measure. In other functional connectivity approaches (I use the term ‘functional connectivity’ in its broadest sense, as a measure that can be made using any functional neuroimaging approach, including EEG, fMRI, MEG, etc.), we correlate the timeseries of activity within subjects to estimate a connectome, whereas with PET the standard was an across-subject approach. So, we thought we’d try to use fPET to estimate a timeseries to correlate, in a similar way. We did this, and published what we think is the first ‘metabolic’ connectome that is estimated using within-subject methodology. The work was hard but rewarding, and we still have lots to learn about how to best process this data, and interpret it.
And it’s this interpretation that’s made me put my hand up and declare myself a diehard convert to molecular connectivity. I’ve focused exclusively on using FDG as a radiotracer, primarily because it’s easy to source, but also because of my early focus on neuronal activity. However, if you’re interested in human physiology, how can you go past the target that literally fuels the brain and body?! My group has subsequently shifted to focus on how individual differences in systemic glucose metabolic efficiency (insulin resistance) influences brain function, the metabolic connectome, and cognition. We’ve got some new results currently under review and about to be submitted that we’re really excited to share with the community in this space.
What is your role in the Molecular Connectivity Working Group and what have you been contributing to/working on with the Molecular Connectivity Working Group?
I’m involved as a Board Member, as well as a member for the symposium council. I’m excited at the opportunity to bring together international interdisciplinary experts in connectivity, to discuss how we can ‘speak the same language’ between the molecular connectivity and other functional connectivity disciplines.
In what ways do you imagine the impact of molecular connectivity will have in advancing our understanding of brain function?
I am really enthusiastic about the possibilities in this space to make important discoveries about brain function. To date, I don’t think the human brain imaging community has really capitalised on the strengths of PET and other modalities to measure molecular targets in vivo in the brain. As such, we have a narrow concept of brain function that we’ve derived from electrophysiology and haemodynamic approaches, that do not adequately take into account the range of molecular targets that we know are important for brain function. As a multimodal neuroimager, I’ve always been inspired by that famous cartoon of the man looking under the streetlamp for his keys. When a passer-by asks him where he lost his keys, he points to a place outside the light and says, ‘Over there, but the light is best over here’. We’re almost always using tools to answer questions that they’re not quite suited to answer. I like to imagine that each neuroimaging method sheds a little more light on the situation: the beams will probably overlap at some level, but also show us something new too. I’m hoping that with the development of the fPET approach, and a renewed focus on molecular connectivity, we will be able to shed new light on human brain function that we couldn’t have achieved with the other methods we have at our disposal.
One aspect that I am excited about, is the ability to use fPET and molecular connectivity with a range of radiotracers to measure dynamic brain activity. During my PhD, I was reading a lot about the role of different neurotransmitters in different cognitive processes, however, these ‘models’ were poorly specified, and had no direct measurements of the neurotransmitter targets to support them. With fPET and molecular connectivity, we can directly measure these targets and test these hypotheses. I think that these methodological advances together with the large improvements in PET detector technology means that we will be seeing a lot of new discoveries in the coming years about the molecular bases of cognition.

The MCWG is made up of four international and multidisciplinary councils dedicated to promoting molecular connectivity research via dissemination of methods, results, collaboration, and resource sharing (e.g. datasets, tools) within the scientific community. We encourage the neuroscientific community to take an integrative perspective in study of the brain connectome, where various methods including MRI-based techniques, electrophysiological tools, and molecular imaging advance our understanding of the brain. Please find fundamental questions outlined here: “Brain connectomics: time for a molecular imaging perspective?”
Our website can be found here. We also invite you to join the MCWG!
[MCWG] Molecular Connectivity Newsletter: February 2024

Hello!
We hope that you enjoyed our inaugural newsletter and wish to thank those that attended our first MCOS in January with Arianna Sala, PhD. We think it was a rousing success and appreciate the participation of everyone who was able to join! We welcome any feedback you may have! Please find the replay here and slides here. We hope to see you at our next talk for further discussion, please find the details below!
1. We are pleased to inform you that the second Molecular Connectivity Working Group Symposium is set for May 3-4 in Munich, Germany
Focusing on brain connectivity, the event features experts in molecular imaging, MRI and EEG. A dedicated session on standardized nomenclature will be exclusively for MCWG members and invited experts.
The symposium will take place in a hybrid format, the preliminary programme is attached: Preliminary Program MCWG Symposium, May 3-4, Munich, Germany.
Website and Registration (opening soon) can be found here. Please share with your colleagues, trainees, collaborators and on social media.
We look forward to seeing you in Munich!
2. We are also very pleased to announce the second talk in our Molecular Connectivity Online Series, ‘MCOS’, featuring Christian Habeck, PhD!
Date: February 23rd, 2024
Time: 15:00 CET, 9:00 EST, 22:00 CST
Title: Basic introduction to multivariate neuroimaging analysis – for nerds and novices
Please join us for this 30 minute presentation to be followed by discussion (~25 minutes).
Please register here.
Abstract: I will (1) go through some of the basics of PCA with toy simulations, (2) discuss some of the recent criticisms of PCA, and (3) show examples in fMRI data to demonstrate the power of multivariate compared to univariate analysis. I hope to convince the audience that an intelligent use of PCA is the best go-to initial pass at complex data, and also provides a good benchmark for more complicated and deeper learning architectures.
Some of the material in this presentation in late 2022: Multivariate analysis (PCA-SSM) of brain data: basic introduction and applications

I was originally trained as a Particle Physicist, but followed the great migration from Physics to the lifesciences after my PhD. (Date undisclosed!). After a brief stint of biophysical computational modeling at the Neurosciences Institute in San Diego, I came to brain imaging analytics with PET and fMRI at Columbia University where I have been ever since. I have conducted, developed, and popularized multivariate analysis frameworks and non-parametric statistics, relying on the trusty workhorse of PCA. My hope is that every practitioner -from seasoned analyst to novice- will get something out of my talk.
The MCOS will include seminars and tutorials to promote resource sharing and methodological rigor in the field. We will hold MCOS every third Friday of the month, subject to change due to speaker availability.
The next MCOS Seminar will be held on March 15th at 15:00 CET, 9:00am EST featuring Vince Calhoun. The title of his talk will be: “NeuroMark PET: Towards a fully automated PET ICA pipeline”. Stay tuned for details in next month’s newsletter!
Members of the MCWG are co-organizing the 12th European Conference on Clinical Neuroimaging in March in Leipzig, featuring invited talks on molecular imaging, total-body PET, imaging in psychiatry and ATN concept in Alzheimer’s disease. Register now to benefit from the Early-Bird-Rates (before Feb 22nd).
Please send us information about events related to brain connectivity and molecular connectivity that you wish to share with the community for consideration no later than the final day of each month using this form.
Please submit any news, including publications you would like highlighted or job listings related to brain connectivity and molecular connectivity that you would like to share with the community for consideration no later than the final day of each month using this form.
Each month, we will feature a member of the MCWG and have a brief Q&A!
This month please enjoy our highlight of Dr. Silvia Paola Caminiti, member of the MCWG Steering Committee.

Dr. Silvia Paola Caminiti is an Assistant Professor at the Department of Brain and Behavioral Sciences at the University of Pavia in Italy. With a background in neuroimaging and neuroscience, Dr. Caminiti is dedicated to advancing our understanding of the intricacies of the brain and its functioning. Dr. Caminiti’s primary research focus revolves around aging and neurodegenerative diseases. The goal of her research program is to examine selective neuronal vulnerability in Alzheimer’s (AD) and Parkinson’s disease (PD) using multimodal neuroimaging including positron emission tomography (PET) and magnetic resonance imaging (MRI). One of her projects aims to evaluate the dynamical connectivity changes affecting the course of AD. By employing advanced technical and computational methods, she passionately seeks to uncover the vulnerabilities of the brain, providing valuable insights into diverse clinical phenotypes.
Dr. Caminiti has graciously responded to our feature questionnaire below!
What is your role in the Molecular Connectivity Working Group and what have you been contributing to/working on with the Molecular Connectivity Working Group?
I’m currently a member of the steering committee for the Molecular Connectivity Working Group, currently serving as the Chair of the Symposium Council. My primary responsibility involves organizing – with the invaluable support of the other Council members – the second edition of the MCWG Symposium titled “Molecular Imaging of Brain Connectivity: Towards Standardized Nomenclature.” This event will be held in Munich on the 3rd and 4th of May 2024. I trust that all of you will thoroughly enjoy the event!
In what ways do you imagine the impact of molecular connectivity will have in advancing our understanding of brain function?
Brain connectivity can be estimated in different ways using PET data. Molecular connectivity provides insights into the intricate molecular mechanisms underlying neuronal processes. For example, molecular imaging allows us to specifically study chemical synapses, which are the predominant mode of signaling in the human brain. The study of brain communication at the molecular level is therefore a necessary step in advancing our understanding of brain connectomics. Moreover, the development of new tracers targeting specific biological substrates could provide further insights into the mechanisms underlying brain connectivity.
What do you think are the most important challenges in current brain connectivity research / what unsolved/underappreciated issues should the community tackle to advance the field?
In my opinion, the most challenging task in the field of brain connectivity research – in the era where a large amount of multi-modal data have been collected – rigorous pipelines of data analyses are essential. It’s crucial to have consistency in data collection and analysis methodologies to compare results across studies and to integrate across-modality evidence. The reliability and reproducibility of findings in the field can be enhanced by creating standardized protocols based on experimental design, imaging techniques, and data processing.
What scientist or scientific achievement do you most admire?
This is a difficult question because I have different favourite scientists and achievements. However, as I am Italian and a woman, Rita Levi Montalcini was by sure my greatest inspiration. I admire her tenacity and dedication to scientific research. The resilience she showed in the face of difficulties, coupled with her enthusiasm to discover new things with curiosity, without being influenced by preconceived ideas, is, in my opinion, a fundamental quality for a good scientist.

The MCWG is made up of four international and multidisciplinary councils dedicated to promoting molecular connectivity research via dissemination of methods, results, collaboration, and resource sharing (e.g. datasets, tools) within the scientific community. We encourage the neuroscientific community to take an integrative perspective in study of the brain connectome, where various methods including MRI-based techniques, electrophysiological tools, and molecular imaging advance our understanding of the brain. Please find fundamental questions outlined here: “Brain connectomics: time for a molecular imaging perspective?”
Our website can be found here. We also invite you to join the MCWG!
[MCWG] Molecular Connectivity Newsletter: January 2024

Hello!
Happy New Year! We are happy to share this inaugural newsletter from the Molecular Connectivity Working Group (MCWG)! We are reaching out to you as you have previously shown interest in the MCWG and we hope that this will be a welcome resource for keeping you informed about future activities and initiatives!
1. We are very pleased to announce the inaugural talk in our Molecular Connectivity Online Series, “MCOS”!
Date: January 19th, 2024
Time: 15:00 CET, 9:00 EST
Title: Brain connectomics: Time for a molecular imaging perspective?
The seminar will comprise a 30 minute presentation followed by discussion (~25 minutes).
Please register here.

Our first talk will feature Arianna Sala, PhD, from Liège University in Belgium, discussing the basics of molecular connectivity. The paper can be found here: “Brain connectomics: time for a molecular imaging perspective?” Arianna’s work aims to better understand disorders of consciousness using a multi-level molecular neuroimaging assessment to investigate patterns of hypometabolism and brain (dis)connectivity.
The MCOS will include seminars and tutorials to promote resource sharing and methodological rigor in the field. We will hold MCOS every third Friday of the month, subject to change due to speaker availability.
Case in point, our seminar for February will be held on 23 February at 15:00 CET, 9:00am EST featuring Chris Habeck, PhD who will discuss Basic introduction to multivariate neuroimaging analysis – for nerds and novices. Stay tuned for next month’s newsletter!
2. New Logo!

MCWG would like to also announce our new logo, created by Michael Rullmann, PhD, member of the Outreach Council. The curved, fresh and almost symmetrical MCWG wordmark echoes the iconic brain illustration of the MCWG cover image in Trends in Cognitive Sciences.
Members of the MCWG are (co-)organizing the 12th European Conference on Clinical Neuroimaging in March in Leipzig, featuring invited talks on molecular imaging, total-body PET, imaging in psychiatry and ATN concept in Alzheimer’s disease. Register now to benefit from the Early-Bird-Rates (before Feb 22nd).
Please send us information about events related to brain connectivity and molecular connectivity that you wish to share with the community for consideration no later than the final day of each month using this form.
Additionally, we would like to share news that MCWG members Silvia Paola Caminiti, Gabriel Gonzales-Escamilla and Arianna Sala served as editors for a recently published Frontiers special issue with papers from additional MCWG members in relation to molecular connectivity found here: Imaging Brain Molecular Connectivity in Health and Disease | Frontiers Research Topic; Editorial: Imaging brain molecular connectivity in health and disease.
Please submit any news, including publications you would like highlighted or job listings related to brain connectivity and molecular connectivity that you would like to share with the community for consideration no later than the final day of each month using this form.
Each month, we will feature a member of the MCWG and have a brief Q&A!
This month please enjoy our highlight of Dr. Simon Eickhoff, member of the MCWG Advisory Board.


Simon Eickhoff is a full professor and chair of the Institute for Systems Neuroscience at the Heinrich-Heine University in Düsseldorf and the director of the Institute of Neuroscience and Medicine (INM-7, Brain and Behavior) at the Forschungszentrum Jülich. He is furthermore a visiting professor at the Chinese Academy of Science Institute of Automation. Working at the interface between neuroanatomy, data-science and brain medicine, he aims to obtain a more detailed characterization of the organization of the human brain and its inter-individual variability in order to better understand its changes in advanced age as well as neurological and psychiatric disorders. This goal is pursued by the development and application of novel analysis tools and approaches for large-scale, multi-modal analysis of brain structure, function and connectivity as well as by machine-learning for single subject prediction of cognitive and socio-affective traits and ultimately precision medicine.
Dr. Eickhoff has graciously responded to our feature questionnaire:
The following three questions were answered in combination.
What is your role in the Molecular Connectivity Working Group and what have you been contributing to/working on with the Molecular Connectivity Working Group?
What do you think are the most important challenges in current brain connectivity research and what unsolved/underappreciated issues should the community tackle to advance the field?
In what ways do you imagine the impact of molecular connectivity will have in advancing our understanding of brain function?
For me personally, these questions are all very closely related. There is no such thing as “the” connectivity between brain areas or “the connectome” of the brain. Rather, each method for assessing brain connectivity can reveal distinct aspects of the brain’s network architecture, its inter-individual variability and ultimately relation to neurological and psychiatric disorders. Unfortunately, each method also comes with its inherent limitations and biases. Consequently, I am deeply convinced that the integration, comparison and combination of different aspects of brain connectivity is the key to get better insights into neurobiology and pathophysiology. And molecular connectivity certainly provides a very distinct layer of information that can provide extremely valuable insight into mechanisms not accessible through any other method. Looking into the respective relationships in health and disease and ultimately using multi-modal information for understanding and predicting behavioural or clinical phenotypes will be the main focus of my work within the MCWG and beyond.
What is your favorite mentoring memory? This can be in relation to the impact of a mentor on you or a memorable story about your impact on a mentee…
It’s not so much a specific moment or event, but what I am most happy about is that we as an institute not only managed the Covid-pandemic but actually developed a new way of working and communicating during that time combining massive flexibility with close interaction. Mentoring people, including faculty, through this process and seeing how nice it works every day is amazing. But observing that it allows many young parents to combine work and family life is my favourite.
What scientist or scientific achievement do you most admire?
Hard to pick any individual one among the pioneers in (clinical) neuroscience during the late 19th century. It is simply unbelievable how much of their basic concepts, from brain organization to clinical nosology is still foundational to our current work, in particular considering the limited methodologies at their disposal. Teaches you a lot about the value of careful investigation and critical conceptualization.
Representative Publications

The MCWG is made up of four international and multidisciplinary councils dedicated to promoting molecular connectivity research via dissemination of methods, results, collaboration, and resource sharing (e.g. datasets, tools) within the scientific community. We encourage the neuroscientific community to take an integrative perspective in study of the brain connectome, where various methods including MRI-based techniques, electrophysiological tools, and molecular imaging advance our understanding of the brain. Please find fundamental questions outlined here: “Brain connectomics: time for a molecular imaging perspective?”